click a reference to see abstract
2023
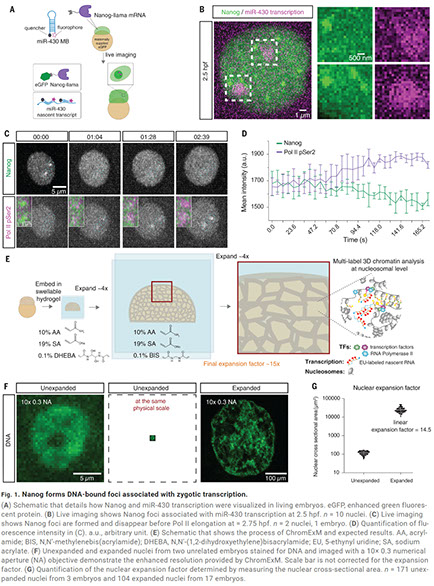
Mark E Pownall, Liyun Miao, Charles E Vejnar, Ons M’Saad, Alice Sherrard, Megan A Frederick, Maria DJ Benitez, Curtis W Boswell, Kenneth S Zaret, Joerg Bewersdorf, Antonio J Giraldez
Chromatin expansion microscopy reveals nanoscale organization of transcription and chromatin
Science 381 (6653), 92-100 https://www.science.org/doi/10.1126/science.ade5308
Nanoscale chromatin organization regulates gene expression. Although chromatin is notably reprogrammed during zygotic genome activation (ZGA), the organization of chromatin regulatory factors during this universal process remains unclear. In this work, we developed chromatin expansion microscopy (ChromExM) to visualize chromatin, transcription, and transcription factors in vivo. ChromExM of embryos during ZGA revealed how the pioneer factor Nanog interacts with nucleosomes and RNA polymerase II (Pol II), providing direct visualization of transcriptional elongation as string-like nanostructures. Blocking elongation led to more Pol II particles clustered around Nanog, with Pol II stalled at promoters and Nanog-bound enhancers. This led to a new model termed "kiss and kick", in which enhancer–promoter contacts are transient and released by transcriptional elongation. Our results demonstrate that ChromExM is broadly applicable to study nanoscale nuclear organization.
Full text
Mary Lou P Bailey, Susan E Pratt, Michael Hinrichsen, Yongdeng Zhang, Joerg Bewersdorf, Lynne J Regan, Simon GJ Mochrie
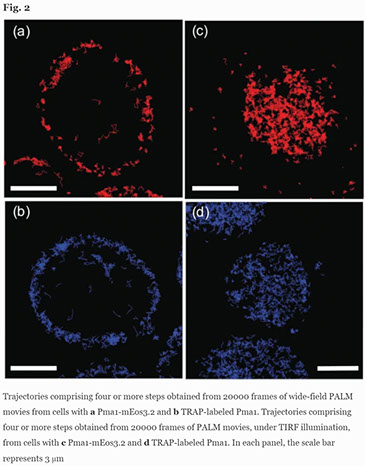
Uncovering diffusive states of the yeast membrane protein, Pma1, and how labeling method can change diffusive behavior
The European Physical Journal E 46 (6), 1-16 https://link.springer.com/article/10.1140/epje/s10189-023-00301-x
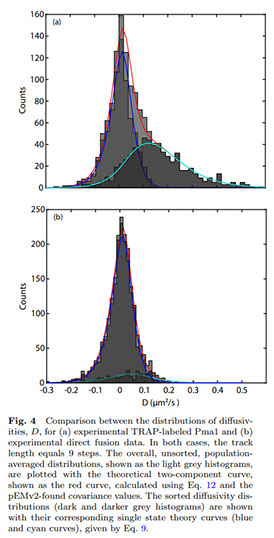
We present and analyze video-microscopy-based single-particle-tracking measurements of the budding yeast (Saccharomyces cerevisiae) membrane protein, Pma1, fluorescently labeled either by direct fusion to the switchable fluorescent protein, mEos3. 2, or by a novel, light-touch, labeling scheme, in which a 5 amino acid tag is directly fused to the C-terminus of Pma1, which then binds mEos3. 2. The track diffusivity distributions of these two populations of single-particle tracks differ significantly, demonstrating that labeling method can be an important determinant of diffusive behavior. We also applied perturbation expectation maximization (pEMv2)(Koo and Mochrie in Phys Rev E 94 (5): 052412, 2016), which sorts trajectories into the statistically optimum number of diffusive states. For both TRAP-labeled Pma1 and Pma1-mEos3. 2, pEMv2 sorts the tracks into two diffusive states: an essentially immobile state and a more mobile state. However, the mobile fraction of Pma1-mEos3.2 tracks is much smaller (∼0.16) than the mobile fraction of TRAP-labeled Pma1 tracks (∼0.43). In addition, the diffusivity of Pma1-mEos3.2’s mobile state is several times smaller than the diffusivity of TRAP-labeled Pma1’s mobile state. Thus, the two different labeling methods give rise to very different overall diffusive behaviors. To critically assess pEMv2’s performance, we compare the diffusivity and covariance distributions of the experimental pEMv2-sorted populations to corresponding theoretical distributions, assuming that Pma1 displacements realize a Gaussian random process. The experiment–theory comparisons for both the TRAP-labeled Pma1 and Pma1-mEos3.2 reveal good agreement, bolstering the pEMv2 approach.
Full text
Vinit Sheth, Xuxin Chen, Evan M Mettenbrink, Wen Yang, Meredith A Jones, Ons M’Saad, Abigail G Thomas, Rylee S Newport, Emmy Francek, Lin Wang, Alex N Frickenstein, Nathan D Donahue, Alyssa Holden, Nathan F Mjema, Dixy E Green, Paul L DeAngelis, Joerg Bewersdorf, Stefan Wilhelm
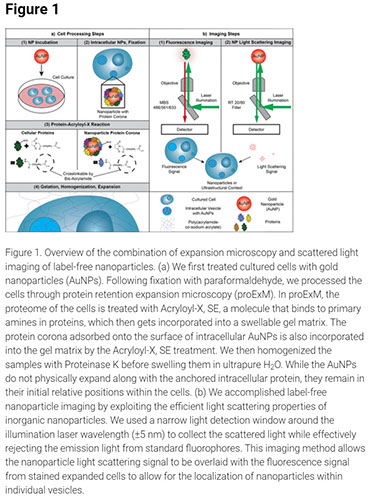
Quantifying Intracellular Nanoparticle Distributions with Three-Dimensional Super-Resolution Microscopy
ACS nano 17 (9), 8376-8392 https://doi.org/10.1021/acsnano.2c12808
Super-resolution microscopy can transform our understanding of nanoparticle–cell interactions. Here, we established a super-resolution imaging technology to visualize nanoparticle distributions inside mammalian cells. The cells were exposed to metallic nanoparticles and then embedded within different swellable hydrogels to enable quantitative three-dimensional (3D) imaging approaching electron-microscopy-like resolution using a standard light microscope. By exploiting the nanoparticles’ light scattering properties, we demonstrated quantitative label-free imaging of intracellular nanoparticles with ultrastructural context. We confirmed the compatibility of two expansion microscopy protocols, protein retention and pan-expansion microscopy, with nanoparticle uptake studies. We validated relative differences between nanoparticle cellular accumulation for various surface modifications using mass spectrometry and determined the intracellular nanoparticle spatial distribution in 3D for entire single cells. This super-resolution imaging platform technology may be broadly used to understand the nanoparticle intracellular fate in fundamental and applied studies to potentially inform the engineering of safer and more effective nanomedicines.
Full text
Andrew ES Barentine, Yu Lin, Edward M Courvan, Phylicia Kidd, Miao Liu, Leonhard Balduf, Timy Phan, Felix Rivera-Molina, Michael R Grace, Zach Marin, Mark Lessard, Juliana Rios Chen, Siyuan Wang, Karla M Neugebauer, Joerg Bewersdorf, David Baddeley
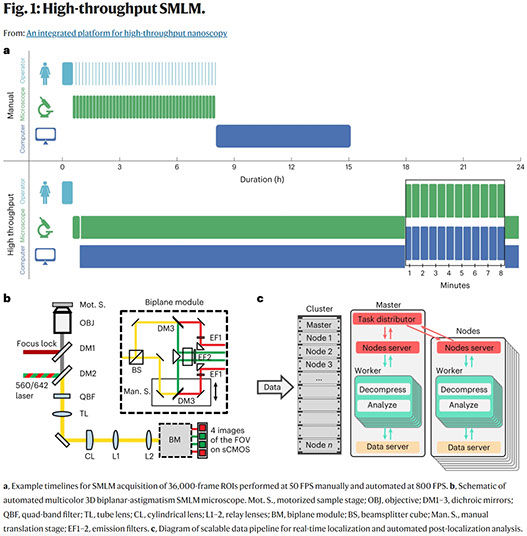
An integrated platform for high-throughput nanoscopy
Nature Biotechnology, 1-8 https://doi.org/10.1038/s41587-023-01702-1
Single-molecule localization microscopy enables three-dimensional fluorescence imaging at tens-of-nanometer resolution, but requires many camera frames to reconstruct a super-resolved image. This limits the typical throughput to tens of cells per day. While frame rates can now be increased by over an order of magnitude, the large data volumes become limiting in existing workflows. Here we present an integrated acquisition and analysis platform leveraging microscopy-specific data compression, distributed storage and distributed analysis to enable an acquisition and analysis throughput of 10,000 cells per day. The platform facilitates graphically reconfigurable analyses to be automatically initiated from the microscope during acquisition and remotely executed, and can even feed back and queue new acquisition tasks on the microscope. We demonstrate the utility of this framework by imaging hundreds of cells per well in multi-well sample formats. Our platform, implemented within the PYthon-Microscopy Environment (PYME), is easily configurable to control custom microscopes, and includes a plugin framework for user-defined extensions.
Full text
Zach Marin, Lukas A Fuentes, Joerg Bewersdorf, David Baddeley
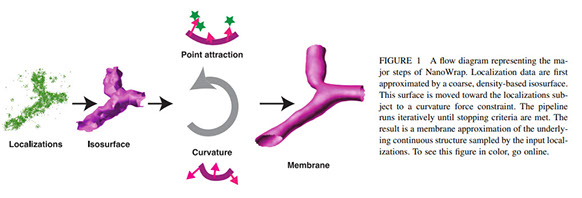
Extracting nanoscale membrane morphology from single-molecule localizations
BioPhysical Journal https://doi.org/10.1016/j.bpj.2023.06.010
Membrane surface reconstruction at the nanometer scale is required for understanding mechanisms of subcellular shape change. This historically has been the domain of electron microscopy, but extraction of surfaces from specific labels is a difficult task in this imaging modality. Existing methods for extracting surfaces from fluorescence microscopy have poor resolution or require high-quality super-resolution data that is manually cleaned and curated. Here we present NanoWrap, a new method for extracting surfaces from generalized single-molecule localization microscopy (SMLM) data. This makes it possible to study the shape of specifically-labelled membraneous structures inside of cells. We validate NanoWrap using simulations and demonstrate its reconstruction capabilities on SMLM data of the endoplasmic reticulum and mitochondria. NanoWrap is implemented in the open-source Python Microscopy Environment.
Full text
Lukas A Fuentes, Zach Marin, Jonathan Tyson, David Baddeley, Joerg Bewersdorf
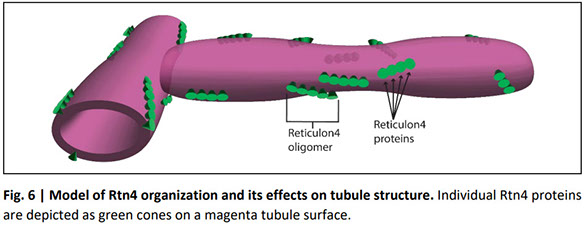
The nanoscale organization of reticulon 4 shapes local endoplasmic reticulum structure in situ
The endoplasmic reticulums (ER) structure is directly linked to the many functions of the ER but its formation is not fully understood. We investigate how the ER-membrane curving protein reticulon 4 (Rtn4) localizes to and organizes in the membrane and how that affects local ER structure. We show a strong correlation between the local Rtn4 density and the local ER membrane curvature. Our data further reveal that the typical ER tubule possesses an elliptical cross-section with Rtn4 enriched at either end of the major axis. Rtn4 oligomers are linear-shaped, contain about five copies of the protein, and preferentially orient parallel to the tubule axis. Our observations support a mechanism in which oligomerization leads to an increase of the local Rtn4 concentration with each molecule increasing membrane curvature through a hairpin wedging mechanism. This quantitative analysis of Rtn4 and its effects on the ER membrane result in a new model of tubule shape as it relates to Rtn4.
Full text
Florian Schueder, Felix Rivera-Molina, Maohan Su, Phylicia Kidd, James E Rothman, Derek Toomre, Joerg Bewersdorf
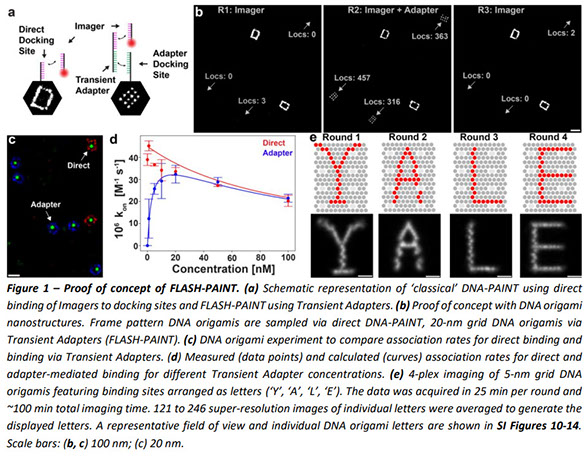
Unraveling cellular complexity with unlimited multiplexed super-resolution imaging
Mapping the intricate spatial relationships between the many different molecules inside a cell is essential to understanding cellular functions in all their complexity. Super-resolution fluorescence microscopy offers the required spatial resolution but struggles to reveal more than four different targets simultaneously. Exchanging labels in subsequent imaging rounds for multiplexed imaging extends this number but is limited by its low throughput. Here we present a novel imaging method for rapid multiplexed super-resolution microscopy of a nearly unlimited number of molecular targets by leveraging fluorogenic labeling in conjunction with Transient Adapter-mediated switching for high-throughput DNA-PAINT (FLASH-PAINT). We demonstrate the cell biological versatility of FLASH-PAINT in mammalian cells in four applications: i) mapping nine proteins in a single mammalian cell, ii) elucidating the functional organization of primary cilia by nine-target imaging, iii) revealing the changes in proximity of twelve different targets in unperturbed and dissociated Golgi stacks and iv) investigating inter-organelle contacts at 3D super-resolution.
Full text
2022
Giulia Biancon, Emma Busarello, Mark D Lessard, Edward Courvan, Dahyana Arias Escayola, Jennifer VanOudenhove, Veronica Lee, Mathew Krick, Joerg Bewersdorf, Karla M Neugebauer, Toma Tebaldi, Stephanie Halene
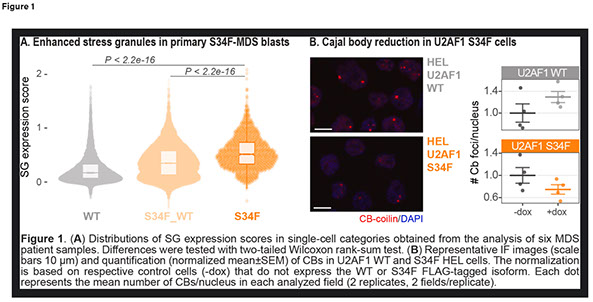
U2AF1 Splicing Factor Mutations Affect RNA Subcellular Compartmentalization in Myeloid Malignancies
Blood 140 (Supplement 1), 1271-1272 https://doi.org/10.1182/blood-2022-170043
Myelodysplastic syndromes (MDS) are a group of clonal hematopoietic disorders characterized by morphological dysplasia, cytopenias and high risk of progression to acute myeloid leukemia (AML). Despite recent improvements in treatment strategies, the 5-year relative survival rate in MDS remains only 31%, dropping to 24% in AML. This highlights the need for a deeper characterization of the pathogenic mechanisms driving disease development and progression.
Full text
Florian Schueder, Joerg Bewersdorf
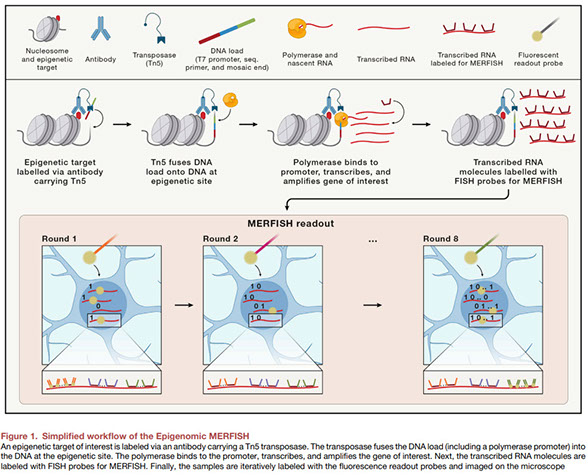
Omics goes spatial epigenomics
Cell 185 (23), 4253-4255 https://doi.org/10.1016/j.cell.2022.10.014
Spatial omics techniques generate spatially resolved, comprehensive data about molecules that define the identity and function of cells in tissues. Epigenetic multiplexing approaches such as Multiplexed Error-robust FISH (MERFISH), introduced by Lu et al.1 in this issue of Cell, now allows researchers to study the epigenomic regulation of gene expression in a tissue-region specific manner.
Full text
Kevin M Dean, Tonmoy Chakraborty, Stephan Daetwyler, Jinlong Lin, Gerard Garrelts, Ons M’Saad, Hannahmariam T Mekbib, Fabian F Voigt, Martina Schaettin, Esther T Stoeckli, Fritjof Helmchen, Joerg Bewersdorf, Reto Fiolka
Isotropic imaging across spatial scales with axially swept light-sheet microscopy
Nature Protocols, 1-29 (2022) https://doi.org/10.1038/s41596-022-00706-6
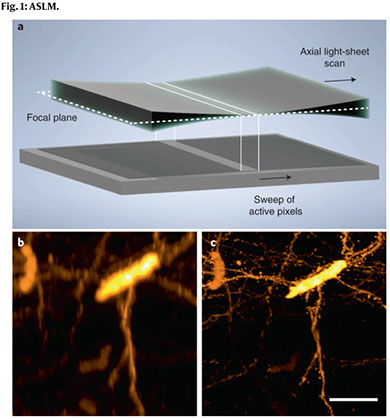
Light-sheet fluorescence microscopy is a rapidly growing technique that has gained tremendous popularity in the life sciences owing to its high-spatiotemporal resolution and gentle, non-phototoxic illumination. In this protocol, we provide detailed directions for the assembly and operation of a versatile light-sheet fluorescence microscopy variant, referred to as axially swept light-sheet microscopy (ASLM), that delivers an unparalleled combination of field of view, optical resolution and optical sectioning. To democratize ASLM, we provide an overview of its working principle and applications to biological imaging, as well as pragmatic tips for the assembly, alignment and control of its optical systems. Furthermore, we provide detailed part lists and schematics for several variants of ASLM that together can resolve molecular detail in chemically expanded samples, subcellular organization in living cells or the anatomical composition of chemically cleared intact organisms. We also provide software for instrument control and discuss how users can tune imaging parameters to accommodate diverse sample types. Thus, this protocol will serve not only as a guide for both introductory and advanced users adopting ASLM, but as a useful resource for any individual interested in deploying custom imaging technology. We expect that building an ASLM will take ~1–2 months, depending on the experience of the instrument builder and the version of the instrument.
Full text
Valentina Greco, Katerina Politi, Stephanie Eisenbarth, Daniel Colón-Ramos, Antonio J Giraldez, Joerg Bewersdorf, David N Berg
A group approach to growing as a principal investigator
Current Biology 32 (11), R498-R504 (2022) https://doi.org/10.1016/j.cub.2022.04.082
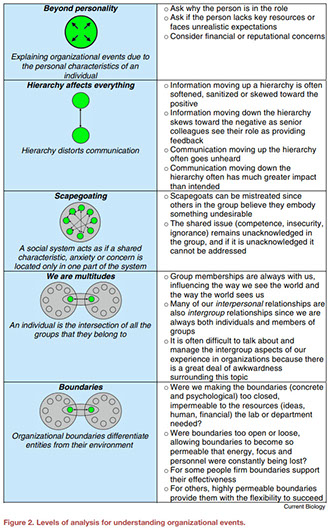
The transition from post-doc to principal investigator (PI) leading an independent academic research laboratory is considered a major milestone for an academic career in the sciences. This transition is accompanied by new roles and responsibilities—beyond directly performing research—for which PIs are generally not trained. These new roles include hiring, managing multiple lab members and having responsibility for the professional performance of their teams. Here, we describe our experiences as early-career professors undergoing this transition. We outline a process we implemented with the guidance of an organizational psychologist to rethink these new roles and responsibilities. We identify common and systemic challenges faced by PIs at academic institutions and describe our decade-long experience in using principles from organizational psychology to help us navigate an academic medical center.
Full text
Kenny KH Chung, Zhao Zhang, Phylicia Kidd, Yongdeng Zhang, Nathan D Williams, Bennett Rollins, Yang Yang, Chenxiang Lin, David Baddeley, Joerg Bewersdorf
Fluorogenic DNA-PAINT for faster, low-background super-resolution imaging
Nature Mehods 19 (5), 554-559 (2022) https://doi.org/10.1038/s41592-022-01464-9
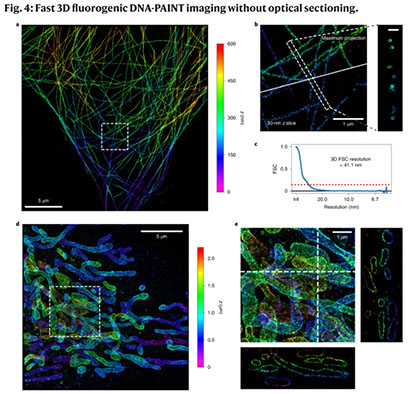
DNA-based points accumulation for imaging in nanoscale topography (DNA-PAINT) is a powerful super-resolution microscopy method that can acquire high-fidelity images at nanometer resolution. It suffers, however, from high background and slow imaging speed, both of which can be attributed to the presence of unbound fluorophores in solution. Here we present two-color fluorogenic DNA-PAINT, which uses improved imager probe and docking strand designs to solve these problems. These self-quenching single-stranded DNA probes are conjugated with a fluorophore and quencher at the terminals, which permits an increase in fluorescence by up to 57-fold upon binding and unquenching. In addition, the engineering of base pair mismatches between the fluorogenic imager probes and docking strands allowed us to achieve both high fluorogenicity and the fast binding kinetics required for fast imaging. We demonstrate a 26-fold increase in imaging speed over regular DNA-PAINT and show that our new implementation enables three-dimensional super-resolution DNA-PAINT imaging without optical sectioning.
Full text
Seong J An, Massimiliano Stagi, Travis J Gould, Yumei Wu, Michael Mlodzianoski, Felix Rivera-Molina, Derek Toomre, Stephen M Strittmatter, Pietro De Camilli, Joerg Bewersdorf, David Zenisek
Multimodal imaging of synaptic vesicles with a single probe
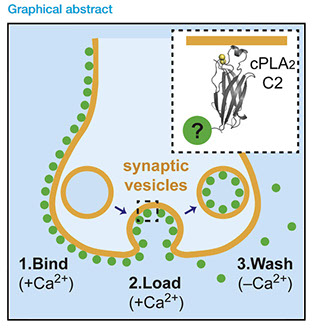
Cell reports methods 2 (4), 100199 (2022) https://doi.org/10.1016/j.crmeth.2022.100199
A complete understanding of synaptic-vesicle recycling requires the use of multiple microscopy methods to obtain complementary information. However, many currently available probes are limited to a specific microscopy modality, which necessitates the use of multiple probes and labeling paradigms. Given the complexity of vesicle populations and recycling pathways, having new single-vesicle probes that could be used for multiple microscopy techniques would complement existing sets of tools for studying vesicle function. Here, we present a probe based on the membrane-binding C2 domain of cytosolic phospholipase A2 (cPLA2) that fulfills this need. By conjugating the C2 domain with different detectable tags, we demonstrate that a single, modular probe can allow synaptic vesicles to be imaged at multiple levels of spatial and temporal resolution. Moreover, as a general endocytic marker, the C2 domain may also be used to study membrane recycling in many cell types.
Giulia Biancon, Poorval Joshi, Joshua T Zimmer, Torben Hunck, Yimeng Gao, Mark D Lessard, Edward Courchaine, Andrew ES Barentine, Martin Machyna, Valentina Botti, Ashley Qin, Rana Gbyli, Amisha Patel, Yuanbin Song, Lea Kiefer, Gabriella Viero, Nils Neuenkirchen, Haifan Lin, Joerg Bewersdorf, Matthew D Simon, Karla M Neugebauer, Toma Tebaldi, Stephanie Halene
Precision analysis of mutant U2AF1 activity reveals deployment of stress granules in myeloid malignancies
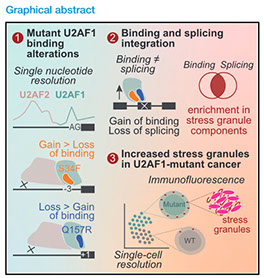
Molecular Cell 82(6), 1107-1122. e7 (2022) https://doi.org/10.1016/j.molcel.2022.02.025
Splicing factor mutations are common among cancers, recently emerging as drivers of myeloid malignancies. U2AF1 carries hotspot mutations in its RNA-binding motifs; however, how they affect splicing and promote cancer remain unclear. The U2AF1/U2AF2 heterodimer is critical for 3′ splice site (3′SS) definition. To specifically unmask changes in U2AF1 function in vivo, we developed a crosslinking and immunoprecipitation procedure that detects contacts between U2AF1 and the 3′SS AG at single-nucleotide resolution. Our data reveal that the U2AF1 S34F and Q157R mutants establish new 3′SS contacts at −3 and +1 nucleotides, respectively. These effects compromise U2AF2-RNA interactions, resulting predominantly in intron retention and exon exclusion. Integrating RNA binding, splicing, and turnover data, we predicted that U2AF1 mutations directly affect stress granule components, which was corroborated by single-cell RNA-seq. Remarkably, U2AF1-mutant cell lines and patient-derived MDS/AML blasts displayed a heightened stress granule response, pointing to a novel role for biomolecular condensates in adaptive oncogenic strategies.
Mary Lou P Bailey, Susan E Pratt, Yongdeng Zhang, Michael Hinrichsen, Joerg Bewersdorf, Lynne J Regan, Simon GJ Mochrie
Uncovering diffusive states of the yeast proton pump, Pma1, and how labeling method can change diffusive behavior
arXiv (2022) https://doi.org/10.48550/arXiv.2201.09358
We present and analyze video-microscopy-based single-particle-tracking measurements of the budding yeast (Saccharomyces cerevisiae) membrane protein, Pma1, fluorescently-labeled either by direct fusion to the switchable fluorescent protein, mEos3.2, or by a novel, light-touch, labeling scheme, in which a 5 amino acid tag is directly fused to the C-terminus of Pma1, which then binds mEos3.2. The diffusivity distributions of these two populations of single particle tracks differ significantly, demonstrating that labeling method can be an important determinant of diffusive behavior. We also applied perturbation expectation maximization (pEMv2) [Physical Review E 94, 052412 (2016)], which sorts trajectories into the statistically-optimum number of diffusive states. For both TRAP-labeled Pma1 and Pma1-mEos3.2, pEMv2 sorts the tracks into two diffusive states: an essentially immobile state and a more mobile state. However, the mobile fraction of Pma1-mEos3.2 tracks is much smaller (0.1) than the mobile fraction of TRAP-labeled Pma1 tracks (0.5). In addition, the diffusivity of Pma1-mEos3.2's mobile state is several times smaller than the diffusivity of TRAP-labeled Pma1's mobile state. To critically assess pEMv2's performance, we compare the diffusivity and covariance distributions of the experimental pEMv2-sorted populations to corresponding theoretical distributions, assuming that Pma1 displacements realize a Gaussian random process. The experiment-theory comparisons for both the TRAP-labeled Pma1 and Pma1-mEos3.2 reveal good agreement, bolstering the pEMv2 approach.
Ons M'Saad, Ravikiran Kasula, Ilona Kondratiuk, Phylicia Kidd, Hanieh Falahati, Juliana E Gentile, Robert F Niescier, Katherine Watters, Robert C Sterner, Seong Lee, Xinran Liu, Pietro De Camilli, James E Rothman, Anthony J Koleske, Thomas Biederer, Joerg Bewersdorf
All-optical visualization of specific molecules in the ultrastructural context of brain tissue
BioRxiv (2022) https://doi.org/10.1101/2022.04.04.486901
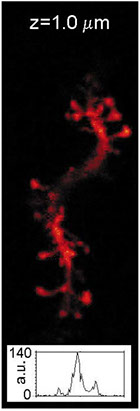
Understanding the molecular anatomy and neural connectivity of the brain requires imaging technologies that can map the 3D nanoscale distribution of specific proteins in the context of brain ultrastructure. Light and electron microscopy (EM) enable visualization of either specific labels or anatomical ultrastructure, but combining molecular specificity with anatomical context is challenging. Here, we present pan-Expansion Microscopy of tissue (pan-ExM-t), an all-optical mouse brain imaging method that combines ∼24-fold linear expansion of biological samples with fluorescent pan-staining of protein densities (providing EM-like ultrastructural context), and immunolabeling of protein targets (for molecular imaging). We demonstrate the versatility of this approach by imaging the established synaptic markers Homer1, Bassoon, PSD-95, Synaptophysin, the astrocytic protein GFAP, myelin basic protein (MBP), and anti-GFP antibodies in dissociated neuron cultures and mouse brain tissue sections. pan-ExM-t reveals these markers in the context of ultrastructural features such as pre and postsynaptic densities, 3D nanoarchitecture of neuropil, and the fine structures of cellular organelles. pan-ExM-t is adoptable in any neurobiological laboratory with access to a confocal microscope and has therefore broad applicability in the research community.
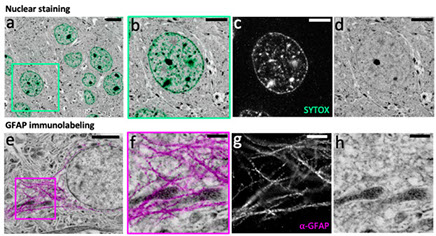
Figure 5: pan-ExM-t is compatible with antibody labeling of brain tissue structural markers.
Full text
2021
Shiwei Zhu, Clinton J Bradfield, Agnieszka Mamińska, Eui-Soon Park, Bae-Hoon Kim, Pradeep Kumar, Shuai Huang, Yongdeng Zhang, Joerg Bewersdorf, John D MacMicking
Cryo-ET of a human GBP coatomer governing cell-autonomous innate immunity to infection
BioRxiv (2021) https://doi.org/10.1101/2021.08.26.457804
All living organisms deploy cell-autonomous defenses to combat infection. In plants and animals, these activities generate large supramolecular complexes that recruit immune proteins for protection. Here, we solve the native structure of a massive antimicrobial complex generated by polymerization of 30,000 human guanylate-binding proteins (GBPs) over the entire surface of virulent bacteria. Construction of this giant nanomachine takes ∼1-3 minutes, remains stable for hours, and acts as a cytokine and cell death signaling platform atop the coated bacterium. Cryo-ET of this “coatomer” revealed thousands of human GBP1 molecules undergo ∼260 Å insertion into the bacterial outer membrane, triggering lipopolysaccharide release that activates co-assembled caspase-4. Together, our results provide a quasi-atomic view of how the GBP coatomer mobilizes cytosolic immunity to combat infection in humans.
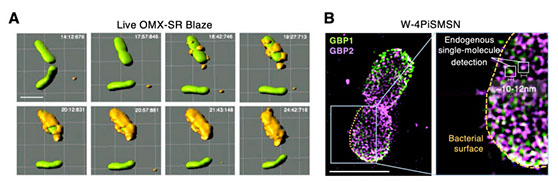
Fig. 1. Human GBP1 coatomer kinetics and functional determinants in situ
Full text
Jonathan Tyson, Kevin Hu, Shuai Zheng, Phylicia Kidd, Neville Dadina, Ling Chu, Derek Toomre, Joerg Bewersdorf, Alanna Schepartz
Extremely bright, near-IR emitting spontaneously blinking fluorophores enable ratiometric multicolor nanoscopy in live cells
ACS central science 7 (8), 1419-1426 (2021) https://opg.optica.org/ao/fulltext.cfm?uri=ao-60-18-5354&id=452022
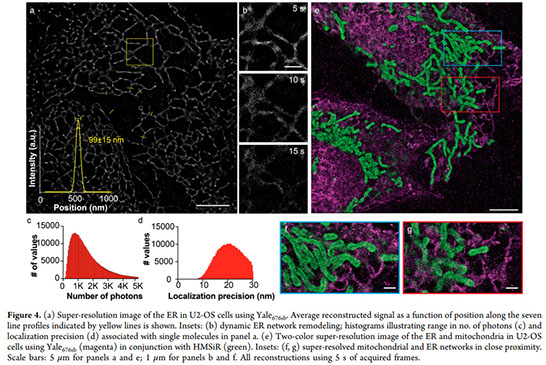
New bright, photostable, emission-orthogonal fluorophores that blink without toxic additives are needed to enable multicolor, live-cell, single-molecule localization microscopy (SMLM). Here we report the design, synthesis, and biological evaluation of Yale676sb, a photostable, near-IR-emitting fluorophore that achieves these goals in the context of an exceptional quantum yield (0.59). When used alongside HMSiR, Yale676sb enables simultaneous, live-cell, two-color SMLM of two intracellular organelles (ER + mitochondria) with only a single laser and no chemical additives.
Full text
Dong-Ryoung Lee and Joerg Bewersdorf
Pupil function design for multifocal confocal, STED, and isoSTED microscopy
Applied Optics Vol. 60, Issue 18, pp. 5354-5359 (2021) https://doi.org/10.1364/AO.416585
Point scanning super-resolution microscopy techniques such as stimulated emission depletion (STED) microscopy are powerful tools to observe biological samples at sub-diffraction limited resolution in three dimensions. However, scanning the sample with only a single beam limits the imaging speed in these microscopes. Here, we propose a concept to increase this speed by introducing highly flexible multifocal illumination and detection. We introduce phase patterns in the objectives’ pupil planes to create arrays of foci in the sample plane with negligible loss of laser power. High uniformity of these foci’s intensities is achieved by iteratively applying a weighted Gerchberg–Saxton phase retrieval algorithm. We characterize the performance of this iterative approach numerically and present simulation results that demonstrate the high quality of the focus arrays for future implementations in laser-scanning STED and isoSTED microscopes. The same approach can also be applied in diffraction-limited confocal laser scanning microscopy.
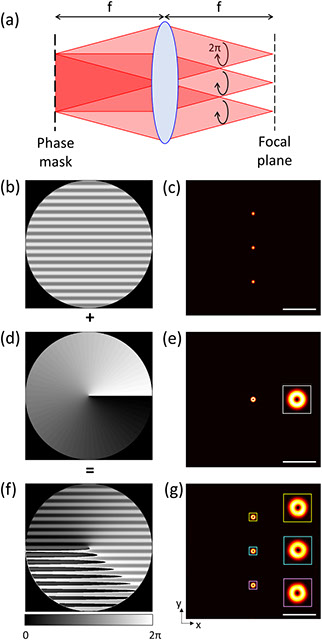
Fig. 6. (a) Geometry of the beam path to generate the multiple doughnut-shaped beams; (b), (d), (f) grating, vortex, and sum patterns and (c), (e), (g) corresponding intensity distributions, respectively. Scale bars, 5 µm.
Full text
Edward M. Courchaine, Andrew E.S. Barentine, Korinna Straube, Dong-Ryoung Lee, Joerg Bewersdorf, Karla M.Neugebauer
DMA-tudor interaction modules control the specificity of in vivo condensates
Cell (2021) https://doi.org/10.1016/j.cell.2021.05.008
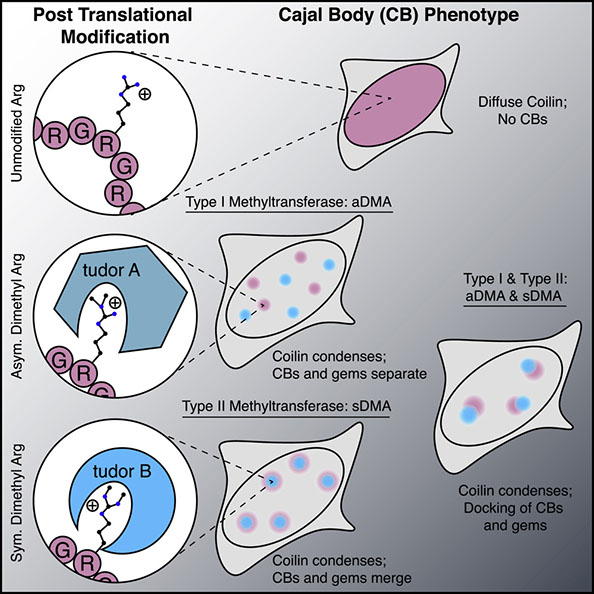
Biomolecular condensation is a widespread mechanism of cellular compartmentalization. Because the “survival of motor neuron protein” (SMN) is implicated in the formation of three different membraneless organelles (MLOs), we hypothesized that SMN promotes condensation. Unexpectedly, we found that SMN’s globular tudor domain was sufficient for dimerization-induced condensation in vivo, whereas its two intrinsically disordered regions (IDRs) were not. Binding to dimethylarginine (DMA) modified protein ligands was required for condensate formation by the tudor domains in SMN and at least seven other fly and human proteins. Remarkably, asymmetric versus symmetric DMA determined whether two distinct nuclear MLOs—gems and Cajal bodies—were separate or “docked” to one another. This substructure depended on the presence of either asymmetric or symmetric DMA as visualized with sub-diffraction microscopy. Thus, DMA-tudor interaction modules—combinations of tudor domains bound to their DMA ligand(s)—represent versatile yet specific regulators of MLO assembly, composition, and morphology.
Full text
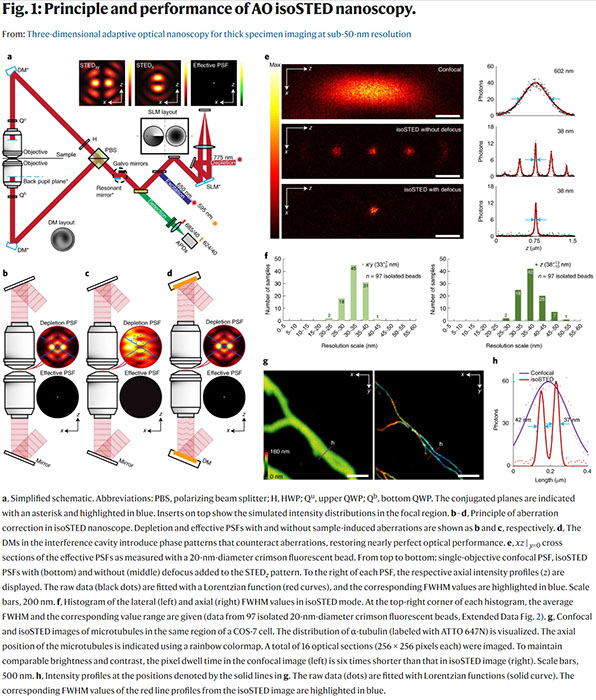
Xiang Hao, Edward S. Allgeyer, Dong-Ryoung Lee, Jacopo Antonello, Katherine Watters, Julianne A. Gerdes, Lena K. Schroeder, Francesca Bottanelli, Jiaxi Zhao, Phylicia Kidd, Mark D. Lessard, James E. Rothman, Lynn Cooley, Thomas Biederer, Martin J. Booth & Joerg Bewersdorf
Three-dimensional adaptive optical nanoscopy for thick specimen imaging at sub-50-nm resolution
Nature Methods (2021) doi: https://doi.org/10.1038/s41592-021-01149-9
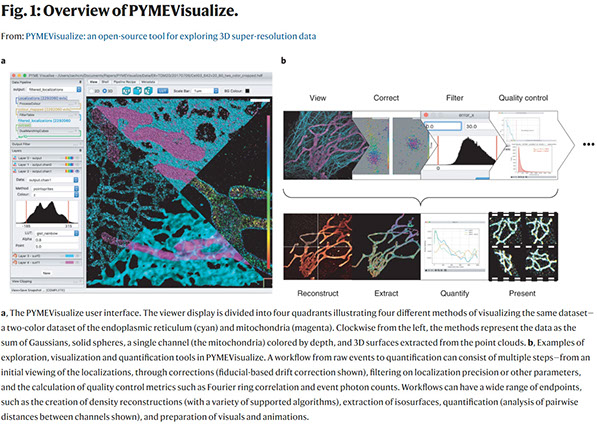
Zach Marin, Michael Graff, Andrew E. S. Barentine, Christian Soeller, Kenny Kwok Hin Chung, Lukas A. Fuentes & David Baddeley
PYMEVisualize: an open-source tool for exploring 3D super-resolution data
Nature Methods (2021) doi: https://doi.org/10.1038/s41592-021-01165-9
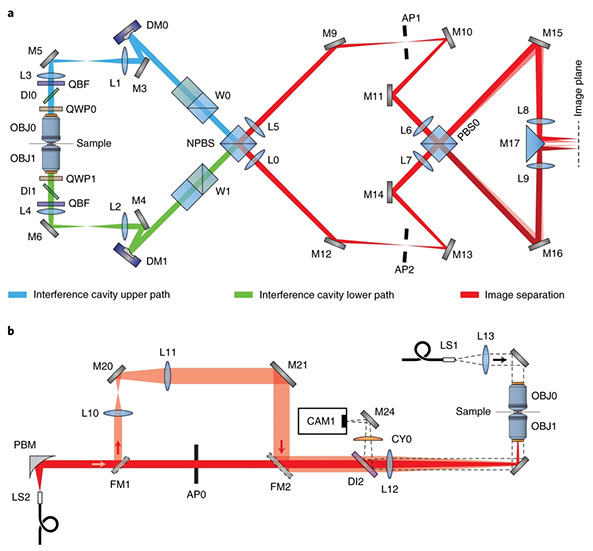
Jingyu Wang, Edward S. Allgeyer, George Sirinakis, Yongdeng Zhang, Kevin Hu, Mark D. Lessard, Yiming Li, Robin Diekmann, Michael A. Phillips, Ian M. Dobbie, Jonas Ries, Martin J. Booth & Joerg Bewersdorf
Implementation of a 4Pi-SMS super-resolution microscope
Nature Protocols (2021) doi:10.1038/s41596-020-00428-7
The development of single-molecule switching (SMS) fluorescence microscopy (also called single-molecule localization microscopy) over the last decade has enabled researchers to image cell biological structures at unprecedented resolution. Using two opposing objectives in a so-called 4Pi geometry doubles the available numerical aperture, and coupling this with interferometric detection has demonstrated 3D resolution down to 10 nm over entire cellular volumes. The aim of this protocol is to enable interested researchers to establish 4Pi-SMS super-resolution microscopy in their laboratories. We describe in detail how to assemble the optomechanical components of a 4Pi-SMS instrument, align its optical beampath and test its performance. The protocol further provides instructions on how to prepare test samples of fluorescent beads, operate this instrument to acquire images of whole cells and analyze the raw image data to reconstruct super-resolution 3D data sets. Furthermore, we provide a troubleshooting guide and present examples of anticipated results. An experienced optical instrument builder will require ~12 months from the start of ordering hardware components to acquiring high-quality biological images.
Full text
N. Bender, M. Sun, H. Yilmaz, J. Bewersdorf, H. Cao
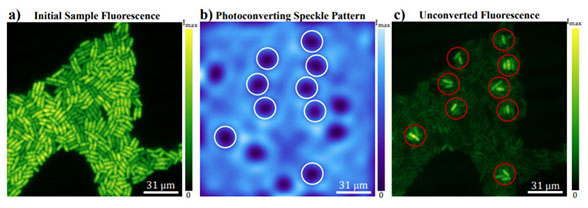
Circumventing the optical diffraction limit with customized speckles
Optica Vol. 8, Issue 2, pp. 122-129 (2021) https://doi.org/10.1364/OPTICA.411007
Speckle patterns have been widely used in imaging techniques such as ghost imaging, dynamic speckle illumination microscopy, structured illumination microscopy, and photoacoustic fluctuation imaging. Recent advances in the ability to control the statistical properties of speckles has enabled the customization of speckle patterns for specific imaging applications. In this work, we design and create special speckle patterns for parallelized nonlinear pattern-illumination microscopy based on fluorescence photoswitching. We present a proof-of-principle experimental demonstration where we obtain a spatial resolution three times higher than the diffraction limit of the illumination optics in our setup. Furthermore, we show that tailored speckles vastly outperform standard speckles. Our work establishes that customized speckles are a potent tool in parallelized super-resolution microscopy.
Full text
M. Sun, K. Hu, J. Bewersdorf*, T.D. Pollard*
Sample Preparation and Imaging Conditions Affect mEos3.2 Photophysics in Fission Yeast
Biophysical Journal Vol. 120, Issue 1, P21-34 (2021) DOI:https://doi.org/10.1016/j.bpj.2020.11.006
*co-corresponding
Photoconvertible fluorescent proteins (PCFPs) are widely used in super-resolution microscopy and studies of cellular dynamics. However, our understanding of their photophysics is still limited, hampering their quantitative application. For example, we do not know the optimal sample preparation methods or imaging conditions to count protein molecules fused to PCFPs by single-molecule localization microscopy in live and fixed cells. We also do not know how the behavior of PCFPs in live cells compares with fixed cells. Therefore, we investigated how formaldehyde fixation influences the photophysical properties of the popular green-to-red PCFP mEos3.2 in fission yeast cells under a wide range of imaging conditions. We estimated photophysical parameters by fitting a 3-state model of photoconversion and photobleaching to the time course of fluorescence signal per yeast cell expressing mEos3.2. We discovered that formaldehyde fixation makes the fluorescence signal, photoconversion rate and photobleaching rate of mEos3.2 sensitive to the buffer conditions by permeabilizing the cell membrane. Under some imaging conditions we tested, the time-integrated mEos3.2 signal per cell is similar in live cells and fixed cells imaged in buffer at pH 8.5 with 1 mM DTT as a reducing agent, indicating that light chemical fixation does not destroy mEos3.2 molecules. We also discovered that 405-nm irradiation
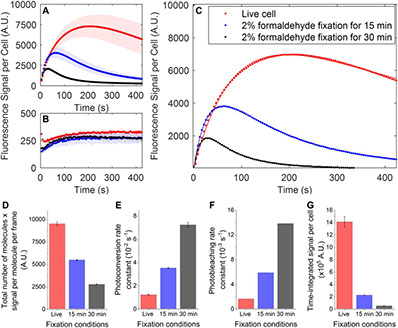
converts some mEos3.2 molecules from the green state to an intermediate state that requires 561-nm illumination for conversion to the red fluorescent state. Our findings provide a guide to compare quantitatively and optimize conditions for imaging and counting of mEos3.2-tagged molecules. Our imaging assay and mathematical model are easy to implement and provide a simple quantitative approach to measure the time-integrated signal and the photoconversion and photobleaching rates of fluorescence proteins in cells.
Full text
M.G.M. Velasco, M. Zhang, J. Antonello, P. Yuan, E.S. Allgeyer, D. May, O. M’Saad, P. Kidd, A.E.S. Barentine, V. Greco, J. Grutzendler, M.J. Booth, J. Bewersdorf
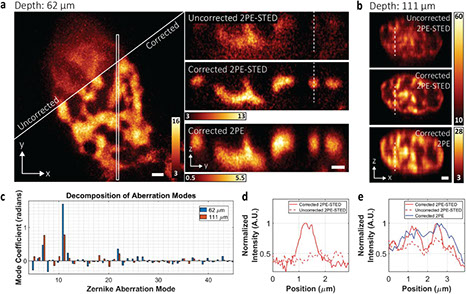
3D super-resolution deep-tissue imaging in living mice
Optica Vol. 8, Issue 4, pp. 442-450 (2021) https://doi.org/10.1364/OPTICA.416841
Stimulated emission depletion (STED) microscopy enables the three-dimensional (3D) visualization of dynamic nanoscale structures in living cells, offering unique insights into their organization. However, 3D-STED imaging deep inside biological tissue is obstructed by optical aberrations and light scattering. We present a STED system that overcomes these challenges. Through the combination of 2-photon excitation, adaptive optics, far-red emitting organic dyes, and a long-working distance water-immersion objective lens, our system achieves aberration-corrected 3D super-resolution imaging, which we demonstrate 164 µm deep in fixed mouse brain tissue and 76 µm deep in the brain of a living mouse.
Full text
2020
L. Tong, P. Yuan, M. Chen, F. Chen, J. Bewersdorf, Z.J. Zhou, J. Grutzendler
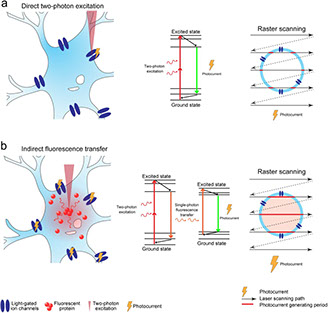
Single cell in vivo brain optogenetic stimulation by two-photon excitation fluorescence transfer
bioRxiv https://doi.org/10.1101/2020.06.29.179077 (2020)
Optogenetics at single-cell resolution can be achieved by two-photon stimulation; however, this requires intense or holographic illumination. We markedly improve stimulation efficiency by positioning fluorophores with high two-photon cross-sections adjacent to opsins. The two-photon-excited fluorescence matches the opsin absorbance and can stimulate opsins in a highly localized manner through efficient single-photon absorption. This indirect fluorescence transfer illumination allows experiments difficult to implement in the live brain such as all-optical neural interrogation and control of regional cerebral blood flow.
Full text
K.K.H. Chung, Z. Zhang, P. Kidd, Y. Zhang, N.D. Williams, B. Rollins, Y. Yang, C. Lin, D. Baddeley, J. Bewersdorf
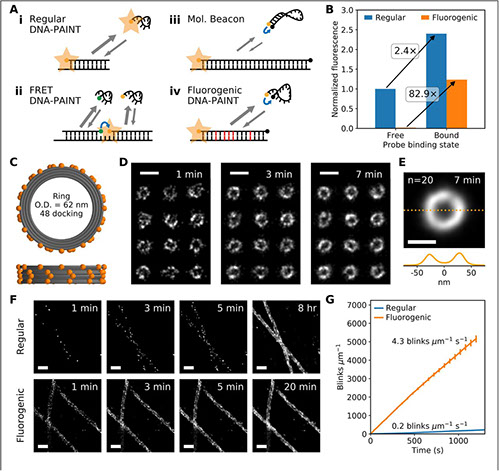
Fluorogenic probe for fast 3D whole-cell DNA-PAINT
bioRxiv, https://doi.org/10.1101/2020.04.29.066886 (2020)
DNA-PAINT is an increasingly popular super-resolution microscopy method that can acquire high-fidelity images at nanometer resolution. It suffers, however, from high background and very slow imaging speed, both of which can be attributed to the presence of unbound fluorophores in solution. We present a fluorogenic DNA-PAINT probe that solves these problems and demonstrate 3D imaging without the need for optical sectioning and a 26-fold increase in imaging speed over regular DNA-PAINT.
Full text
Y. Li, E. Buglakova, Y. Zhang, J.V. Thevathasan, J. Bewersdorf, J. Ries
Accurate 4Pi single-molecule localization using an experimental PSF model
Optics Letters 45(13): 3765-3768. https://doi.org/10.1364/OL.397754 (2020)
Interferometric single-molecule localization microscopy (iPALM, 4Pi-SMS) uses multiphase interferometry to localize single fluorophores and achieves nanometer isotropic resolution in 3D. The current data analysis workflow, however, fails to reach the theoretical resolution limit due to the suboptimal localization algorithm. Here, we develop a method to fit an experimentally derived point spread function (PSF) model to the interference 4Pi-PSF. As the interference phase is not fixed with respect to the shape of the PSF, we decoupled the phase term in the model from the 3D position of the PSF. The fitter can reliably infer the interference period even without introducing astigmatism, reducing the complexity of the microscope. Using a spline-interpolated experimental PSF model and by fitting all phase images globally, we show on simulated data that we can achieve the theoretical limit of 3D resolution, the Cramér–Rao lower bound (CRLB), also for the 4Pi microscope.
Full text
O. M’Saad, J. Bewersdorf
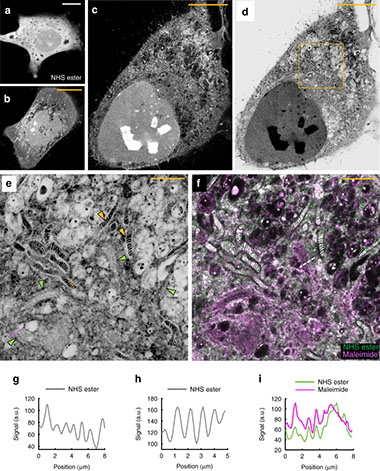
Light microscopy of proteins in their ultrastructural context
Nature Communications 11: 3850. https://doi.org/10.1038/s41467-020-17523-8 (2020)
Resolving the distribution of specific proteins at the nanoscale in the ultrastructural context of the cell is a major challenge in fluorescence microscopy. We report the discovery of a new principle for an optical contrast equivalent to electron microscopy (EM) which reveals the ultrastructural context of the cells with a conventional confocal microscope. By decrowding the intracellular space through 13 to 21-fold physical expansion while simultaneously retaining the proteins, bulk (pan) labeling of the proteome resolves local protein densities and reveals the cellular nanoarchitecture by standard light microscopy.
Full text
Yongdeng Zhang*, Lena K. Schroeder*, Mark D. Lessard, Phylicia Kidd, Jeeyun Chung, Yoanbin Song, Lorena Benedetti, Yiming Li, Jonas Ries, Jonathan B. Grimm, Luke D. Lavis, Pietro De Camilli, James E. Rothman, David Baddeley, Joerg Bewersdorf.
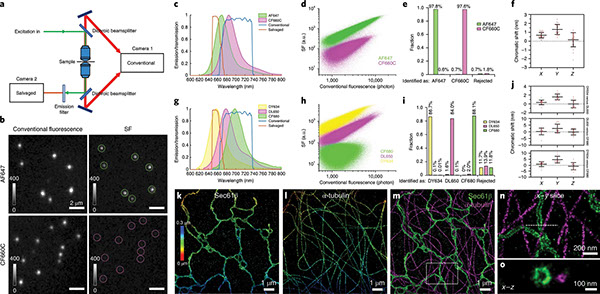
Nanoscale subcellular architecture revealed by multicolor three-dimensional salvaged fluorescence imaging.
Nat Methods (2020) doi:10.1038/s41592-019-0676-4 *equal authorship
Combining the molecular specificity of fluorescent probes with three-dimensional imaging at nanoscale resolution is critical for investigating the spatial organization and interactions of cellular organelles and protein complexes. We present a 4Pi single-molecule switching super-resolution microscope that enables ratiometric multicolor imaging of mammalian cells at 5–10-nm localization precision in three dimensions using ‘salvaged fluorescence’. Imaging two or three fluorophores simultaneously, we show fluorescence images that resolve the highly convoluted Golgi apparatus and the close contacts between the endoplasmic reticulum and the plasma membrane, structures that have traditionally been the imaging realm of electron microscopy. The salvaged fluorescence approach is equally applicable in most single-objective microscopes.
Full text
2019
A.E.S Barentine, Y. Lin, M. Liu, P. Kidd, L. Balduf, M.R. Grace, S. Wang, J. Bewersdorf*, D. Baddeley*
3D Multicolor Nanoscopy at 10,000 Cells a Day
bioRxiv 606954; https://doi.org/10.1101/606954 (2019)
*co-corresponding
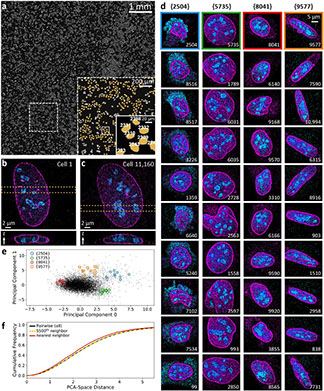
Diffraction-unlimited single-molecule switching (SMS) nanoscopy techniques like STORM /(F)PALM enable three-dimensional (3D) fluorescence imaging at 20-80 nm resolution and are invaluable to investigate sub-cellular organization. They suffer, however, from low throughput, limiting the output of a days worth of imaging to typically a few tens of mammalian cells. Here we develop an SMS imaging platform that combines high-speed 3D single-molecule data acquisition with an automated, fully integrated, high-volume data processing pipeline. We demonstrate 2-color 3D super-resolution imaging of over 10,000 mammalian cell nuclei in about 26 hours, connecting the traditionally low-throughput super-resolution community to the world of omics approaches.
Full text
Hwang JY, Mannowetz N, Zhang Y, Everley RA, Gygi SP, Bewersdorf J, Lishko PV, Chung JJ.
Dual Sensing of Physiologic pH and Calcium by EFCAB9 Regulates Sperm Motility
Cell. 2019 May 30;177(6):1480-1494.e19. doi: 10.1016/j.cell.2019.03.047. Epub 2019 May 2
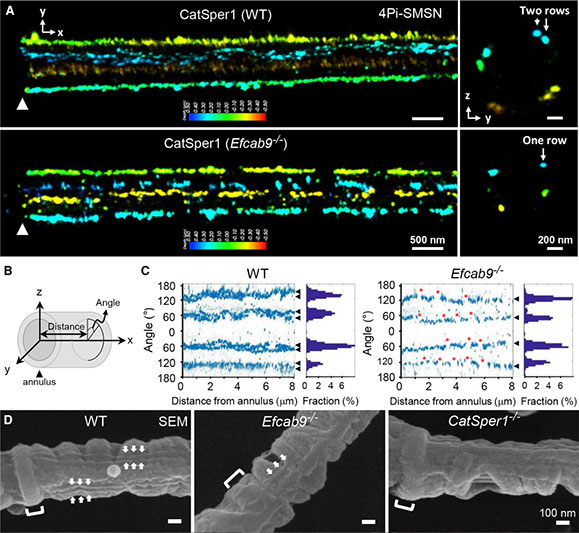
Varying pH of luminal fluid along the female reproductive tract is a physiological cue that modulates sperm motility. CatSper is a sperm-specific, pH-sensitive calcium channel essential for hyperactivated motility and male fertility. Multi-subunit CatSper channel complexes organize linear Ca2+ signaling nanodomains along the sperm tail. Here, we identify EF-hand calcium-binding domain-containing protein 9 (EFCAB9) as a bifunctional, cytoplasmic machine modulating the channel activity and the domain organization of CatSper. Knockout mice studies demonstrate that EFCAB9, in complex with the CatSper subunit, CATSPERζ, is essential for pH-dependent and Ca2+-sensitive activation of the CatSper channel. In the absence of EFCAB9, sperm motility and fertility is compromised, and the linear arrangement of the Ca2+ signaling domains is disrupted. EFCAB9 interacts directly with CATSPERζ in a Ca2+-dependent manner and dissociates at elevated pH. These observations suggest that EFCAB9 is a long-sought, intracellular, pH-dependent Ca2+ sensor that triggers changes in sperm motility.
Full text
Erdmann RS, Baguley SW, Richens JH, Wissner RF, Xi Z, Allgeyer ES, Zhong S, Thompson AD, Lowe N, Butler R, Bewersdorf J, Rothman JE, St Johnston D, Schepartz A, Toomre D.
Labeling Strategies Matter for Super-Resolution Microscopy: A Comparison between HaloTags and SNAP-tags
Cell Chem Biol. 2019 Apr 18;26(4):584-592.e6. doi: 10.1016/j.chembiol.2019.01.003. Epub 2019 Feb 7
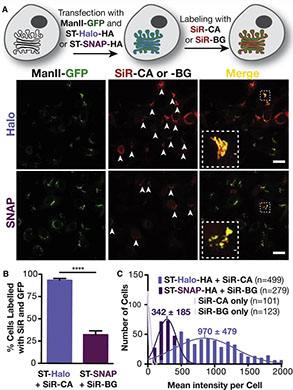
Super-resolution microscopy requires that subcellular structures are labeled with bright and photostable fluorophores, especially for live-cell imaging. Organic fluorophores may help here as they can yield more photons—by orders of magnitude—than fluorescent proteins. To achieve molecular specificity with organic fluorophores in live cells, self-labeling proteins are often used, with HaloTags and SNAP-tags being the most common. However, how these two different tagging systems compare with each other is unclear, especially for stimulated emission depletion (STED) microscopy, which is limited to a small repertoire of fluorophores in living cells. Herein, we compare the two labeling approaches in confocal and STED imaging using various proteins and two model systems. Strikingly, we find that the fluorescent signal can be up to 9-fold higher with HaloTags than with SNAP-tags when using far-red rhodamine derivatives. This result demonstrates that the labeling strategy matters and can greatly influence the duration of super-resolution imaging.
Full text
2018
Lena K. Schroeder, Andrew E.S. Barentine, Holly Merta, Sarah Schweighofer, Yongdeng Zhang, David Baddeley, Joerg Bewersdorf, Shirin Bahmanyar
Dynamic nanoscale morphology of the ER surveyed by STED microscopy
J Cell Biol Nov 2018, jcb.201809107; DOI: 10.1083/jcb.201809107
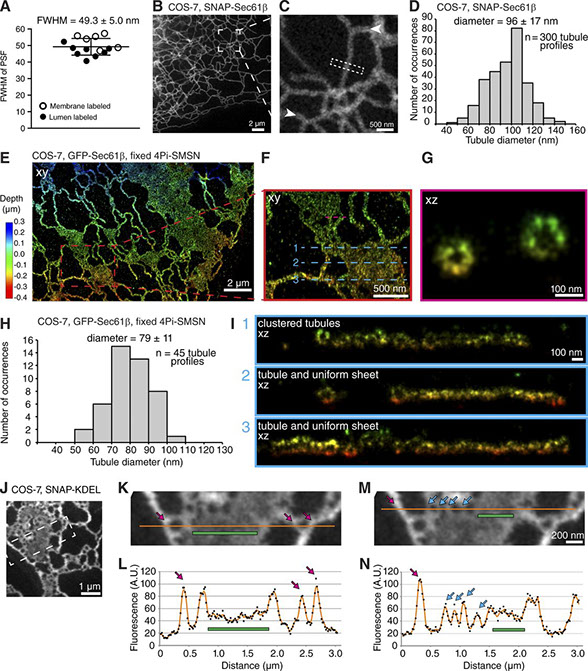
The endoplasmic reticulum (ER) is composed of interconnected membrane sheets and tubules. Superresolution microscopy recently revealed densely packed, rapidly moving ER tubules mistaken for sheets by conventional light microscopy, highlighting the importance of revisiting classical views of ER structure with high spatiotemporal resolution in living cells. In this study, we use live-cell stimulated emission depletion (STED) microscopy to survey the architecture of the ER at 50-nm resolution. We determine the nanoscale dimensions of ER tubules and sheets for the first time in living cells. We demonstrate that ER sheets contain highly dynamic, subdiffraction-sized holes, which we call nanoholes, that coexist with uniform sheet regions. Reticulon family members localize to curved edges of holes within sheets and are required for their formation. The luminal tether Climp63 and microtubule cytoskeleton modulate their nanoscale dynamics and organization. Thus, by providing the first quantitative analysis of ER membrane structure and dynamics at the nanoscale, our work reveals that the ER in living cells is not limited to uniform sheets and tubules; instead, we suggest the ER contains a continuum of membrane structures that includes dynamic nanoholes in sheets as well as clustered tubules.
Full text
Nicole Kilian, Alexander Goryaynov, Mark D. Lessard, Giles Hooker, Derek Toomre, James E. Rothman & Joerg Bewersdorf
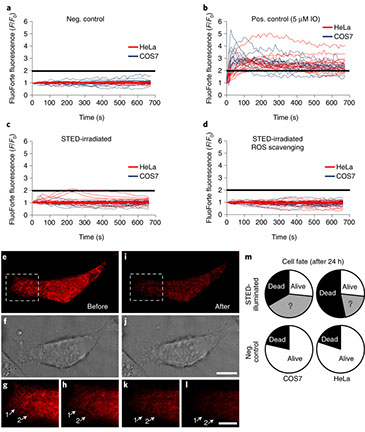
Assessing photodamage in live-cell STED microscopy
Nature Methods, 15: 755-756 (2018)
The recent breakthroughs in the development of optical nanoscopy have provided unprecedented views of the inner workings of cells. Stimulated emission depletion (STED) microscopy, in particular, allows real-time observation of living cells at resolutions of 50 nm or less1,2. However, the high irradiation intensities used in STED nanoscopy have raised concerns about the validity of live-cell observations obtained with this and similar approaches3,4. We report here that, under the right conditions, living cells can be imaged by STED nanoscopy without substantial photodamage.
Andrew E.S. Barentine, Lena K. Schroeder, Michael Graff, David Baddeley, Joerg Bewersdorf
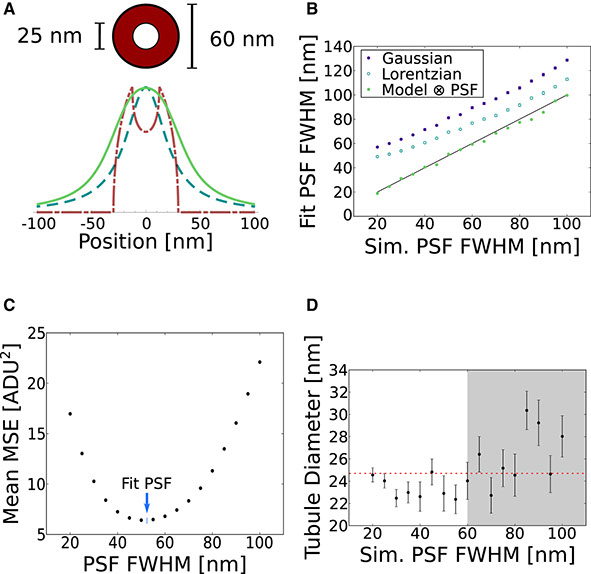
Simultaneously Measuring Image Features and Resolution in Live-Cell STED Images
Biophysical Journal, 115: 951-956 (2018)
Reliable interpretation and quantification of cellular features in fluorescence microscopy requires an accurate estimate of microscope resolution. This is typically obtained by measuring the image of a nonbiological proxy for a point-like object, such as a fluorescent bead. Although appropriate for confocal microscopy, bead-based measurements are problematic for stimulated emission depletion microscopy and similar techniques where the resolution depends critically on the choice of fluorophore and acquisition parameters. In this article, we demonstrate that for a known geometry (e.g., tubules), the resolution can be measured in situ by fitting a model that accounts for both the point spread function (PSF) and the fluorophore distribution. To address the problem of coupling between tubule diameter and PSF width, we developed a technique called nested-loop ensemble PSF fitting. This approach enables extraction of the size of cellular features and the PSF width in fixed-cell and live-cell images without relying on beads or precalibration. Nested-loop ensemble PSF fitting accurately recapitulates microtubule diameter from stimulated emission depletion images and can measure the diameter of endoplasmic reticulum tubules in live COS-7 cells. Our algorithm has been implemented as a plugin for the PYthon Microscopy Environment, a freely available and open-source software.
Full text
Karanastasis AA, Zhang Y, Kenath GS, Lessard MD, Bewersdorf J, Ullal CK.
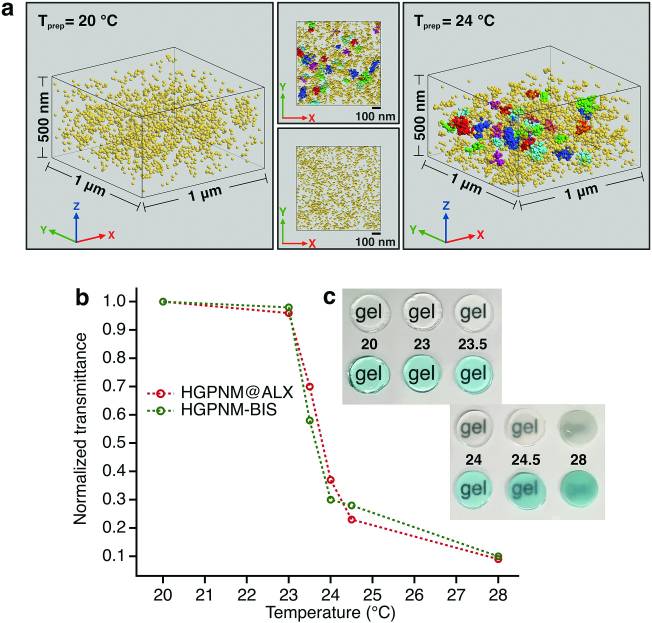
3D mapping of nanoscale crosslink heterogeneities in microgels.
Mater Horiz. 5(6):1130-1136 (2018)
The majority of swollen polymer networks exhibit spatial variations in crosslink density. These spatial heterogeneities are particularly important in colloidal gel particles, or microgels, where they manifest themselves on the nanoscale and impact mechanical and transport properties. Despite their importance, the real space nanostructure of these heterogeneities at the individual particle level has remained elusive. Using state of the art super-resolution microscopy known as Whole cell 4Pi Single Molecule Switching Nanoscopy (W-4PiSMSN) we demonstrate 3D nanoscale mapping of spatial crosslink heterogeneities in a model system of poly(N-isopropylacrylamide) colloidal gel particles containing a novel fluorophore tagged crosslinker. We reveal the presence of higher crosslink density clusters embedded in a lower crosslink density matrix within the core of individual microgel particles, a phenomenon that has been predicted, but never been observed before in real space. The morphology of the clusters provides insight into the kinetics of microgel formation. This study also provides proof-of-concept 3D super-resolution imaging of spatial heterogeneities in bulk hydrogels.
Full text
David Baddeley and Joerg Bewersdorf
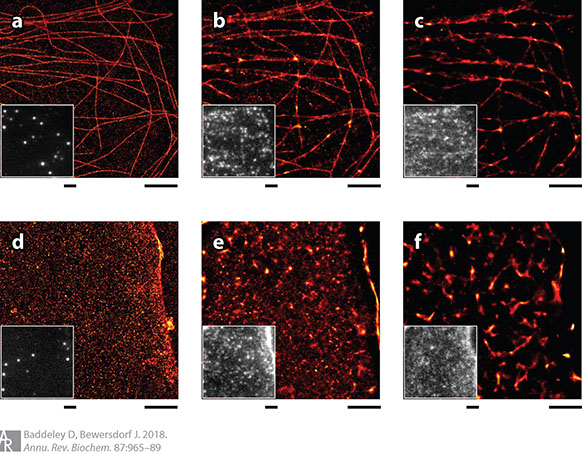
Biological Insight from Super-Resolution Microscopy: What We Can Learn from Localization-Based Images
Annual Review of Biochemistry 87:3.1-3.25 2018.
Super-resolution optical imaging based on the switching and localization of individual fluorescent molecules [photoactivated localization microscopy (PALM), stochastic optical reconstruction microscopy (STORM), etc.] has evolved remarkably over the last decade. Originally driven by pushing technological limits, it has become a tool of biological discovery. The initial demand for impressive pictures showing well-studied biological structures has been replaced by a need for quantitative, reliable data providing dependable evidence for specific unresolved biological hypotheses. In this review, we highlight applications that showcase this development, identify the features that led to their success, and discuss remaining challenges and difficulties. In this context, we consider the complex topic of defining resolution for this imaging modality and address some of the more common analytical methods used with this data.
Full text
Lorena Benedetti, Andrew E. S. Barentine, Mirko Messa, Heather Wheeler, Joerg Bewersdorf and Pietro De Camilli
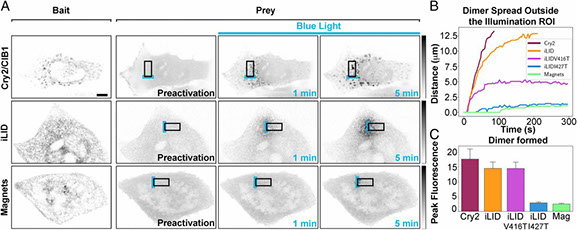
Light-activated protein interaction with high spatial subcellular confinement
PNAS, 115 (10) E2238-E2245; Epub 2018 February 20
Methods to acutely manipulate protein interactions at the subcellular level are powerful tools in cell biology. Several blue-light-dependent optical dimerization tools have been developed. In these systems one protein component of the dimer (the bait) is directed to a specific subcellular location, while the other component (the prey) is fused to the protein of interest. Upon illumination, binding of the prey to the bait results in its subcellular redistribution. Here, we compared and quantified the extent of light-dependent dimer occurrence in small, subcellular volumes controlled by three such tools: Cry2/CIB1, iLID, and Magnets. We show that both the location of the photoreceptor protein(s) in the dimer pair and its (their) switch-off kinetics determine the subcellular volume where dimer formation occurs and the amount of protein recruited in the illuminated volume. Efficient spatial confinement of dimer to the area of illumination is achieved when the photosensitive component of the dimerization pair is tethered to the membrane of intracellular compartments and when on and off kinetics are extremely fast, as achieved with iLID or Magnets. Magnets and the iLID variants with the fastest switch-off kinetics induce and maintain protein dimerization in the smallest volume, although this comes at the expense of the total amount of dimer. These findings highlight the distinct features of different optical dimerization systems and will be useful guides in the choice of tools for specific applications.
Full text
2017
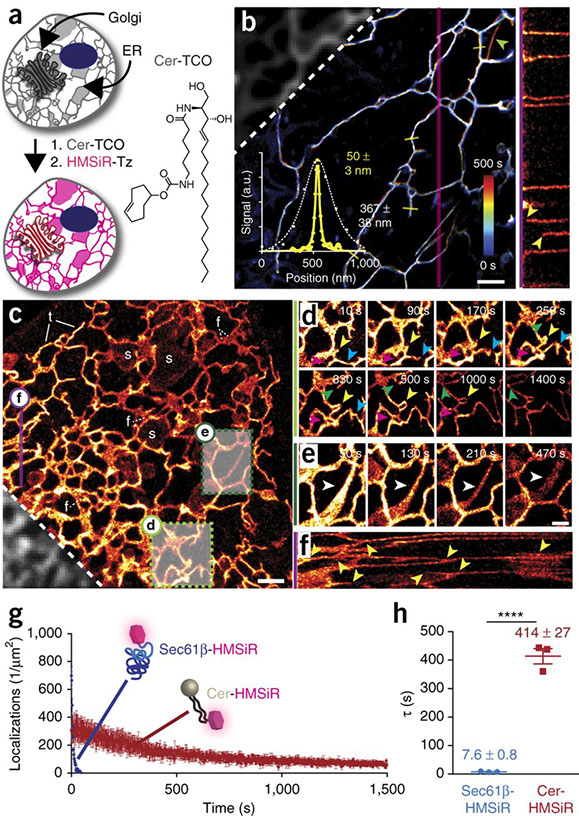
Hideo Takakura, Yongdeng Zhang, Roman S Erdmann, Alexander D Thompson, Yu Lin, Brian McNellis, Felix Rivera-Molina, Shin-nosuke Uno, Mako Kamiya, Yasuteru Urano, James E Rothman, Joerg Bewersdorf, Alanna Schepartz & Derek Toomre
Long time-lapse nanoscopy with spontaneously blinking membrane probes
Nature Biotechnology, doi:10.1038/nbt.3876 Epub 2017 July 03
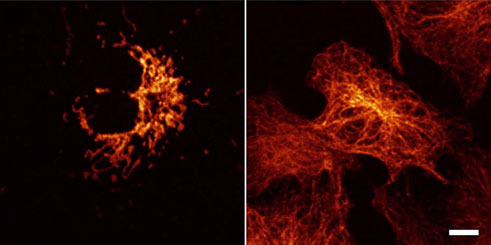
Imaging cellular structures and organelles in living cells by long time-lapse super-resolution microscopy is challenging, as it requires dense labeling, bright and highly photostable dyes, and non-toxic conditions. We introduce a set of high-density, environment-sensitive (HIDE) membrane probes, based on the membrane-permeable silicon-rhodamine dye HMSiR, that assemble in situ and enable long time-lapse, live-cell nanoscopy of discrete cellular structures and organelles with high spatiotemporal resolution. HIDE-enabled nanoscopy movies span tens of minutes, whereas movies obtained with labeled proteins span tens of seconds. Our data reveal 2D dynamics of the mitochondria, plasma membrane and filopodia, and the 2D and 3D dynamics of the endoplasmic reticulum, in living cells. HIDE probes also facilitate acquisition of live-cell, two-color, super-resolution images, expanding the utility of nanoscopy to visualize dynamic processes and structures in living cells.
Full text
Bottanelli F, Kilian N, Ernst AM, Rivera-Molina F, Schroeder LK, Kromann EB, Lessard MD, Erdmann RS, Schepartz A, Baddeley D, Bewersdorf J, Toomre D, Rothman JE.
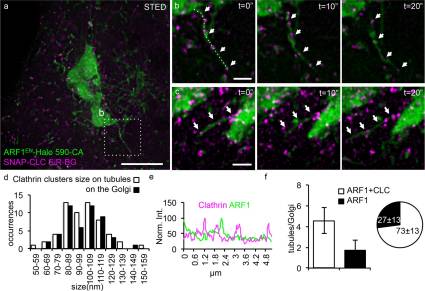
A novel physiological role for ARF1 in the formation of bidirectional tubules from the Golgi.
Mol Biol Cell. 2017 Jun 15;28(12):1676-1687. doi: 10.1091/mbc.E16-12-0863. Epub 2017 Apr 20.
Capitalizing on CRISPR/Cas9 gene-editing techniques and super-resolution nanoscopy, we explore the role of the small GTPase ARF1 in mediating transport steps at the Golgi. Besides its well-established role in generating COPI vesicles, we find that ARF1 is also involved in the formation of long (∼3 µm), thin (∼110 nm diameter) tubular carriers. The anterograde and retrograde tubular carriers are both largely free of the classical Golgi coat proteins coatomer (COPI) and clathrin. Instead, they contain ARF1 along their entire length at a density estimated to be in the range of close packing. Experiments using a mutant form of ARF1 affecting GTP hydrolysis suggest that ARF1[GTP] is functionally required for the tubules to form. Dynamic confocal and stimulated emission depletion imaging shows that ARF1-rich tubular compartments fall into two distinct classes containing 1) anterograde cargoes and clathrin clusters or 2) retrograde cargoes and coatomer clusters.
Full text
Zhang Y^, Lara-Tejero M^, Bewersdorf J*, Galán JE*.
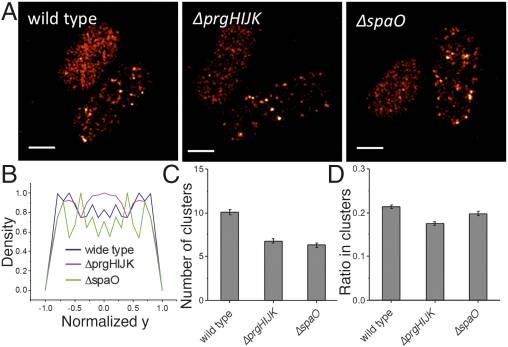
Visualization and characterization of individual type III protein secretion machines in live bacteria.
Proc Natl Acad Sci U S A. 2017 Jun 6;114(23):6098-6103. doi: 10.1073/pnas.1705823114. Epub 2017 May 22.
^ equal authorship
* co-corresponding
Type III protein secretion machines have evolved to deliver bacterially encoded effector proteins into eukaryotic cells. Although electron microscopy has provided a detailed view of these machines in isolation or fixed samples, little is known about their organization in live bacteria. Here we report the visualization and characterization of the Salmonella type III secretion machine in live bacteria by 2D and 3D single-molecule switching superresolution microscopy. This approach provided access to transient components of this machine, which previously could not be analyzed. We determined the subcellular distribution of individual machines, the stoichiometry of the different components of this machine in situ, and the spatial distribution of the substrates of this machine before secretion. Furthermore, by visualizing this machine in Salmonella mutants we obtained major insights into the machine's assembly. This study bridges a major resolution gap in the visualization of this nanomachine and may serve as a paradigm for the examination of other bacterially encoded molecular machines.
Full text
Hao, Xiang; Antonello, Jacopo; Allgeyer, Edward S.; Bewersdorf*, Joerg; Booth*, Martin J.
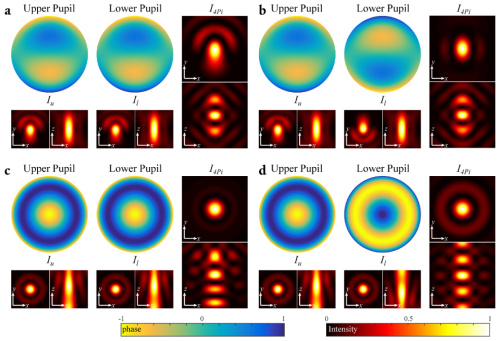
Aberrations in 4Pi Microscopy
Optics Express 25(13) 14049-14058 (2017)
* co-corresponding
The combination of two opposing objective lenses in 4Pi fluorescence microscopy significantly improves the axial resolution and increases the collection efficiency. Combining 4Pi microscopy with other super-resolution techniques has resulted in the highest three-dimensional (3D) resolution in fluorescence microscopy to date. It has previously been shown that the performance of 4Pi microscopy is significantly affected by aberrations. However, a comprehensive description of 4Pi microscope aberrations has been missing. In this paper, we introduce an approach to describe aberrations in a 4Pi cavity through a new functional representation. We discuss the focusing properties of 4Pi systems affected by aberrations and discuss the implications for adaptive optics schemes for 4Pi microscopes based on this new insight.
Full text
Akamatsu M, Lin Y, Bewersdorf J, Pollard TD.
Analysis of interphase node proteins in fission yeast by quantitative and super resolution fluorescence microscopy.
Mol Biol Cell. 2017 May 24. pii: mbc.E16-07-0522. doi: 10.1091/mbc.E16-07-0522. [Epub ahead of print]
We used quantitative confocal microscopy and FPALM super resolution microscopy of live fission yeast to investigate the structures and assembly of two types of interphase nodes, multiprotein complexes associated with the plasma membrane that merge together and mature into the precursors of the cytokinetic contractile ring. During the long G2 phase of the cell cycle seven different interphase node proteins maintain constant concentrations as they accumulate in proportion to cell volume. During mitosis the total numbers of type 1 node proteins (cell cycle kinases Cdr1p, Cdr2p, Wee1p, and anillin Mid1p) are constant even when the nodes disassemble. Quantitative measurements provide strong evidence that both types of nodes have defined sizes and numbers of constituent proteins, as observed for cytokinesis nodes. Type 1 nodes assemble in two phases, a burst at the end of mitosis, followed by steady increase during interphase to double the initial number. Type 2 nodes containing Blt1p, Rho-GEF Gef2p, and kinesin Klp8p remain intact throughout the cell cycle and are constituents of the contractile ring. They are released from the contractile ring as it disassembles and then associate with type 1 nodes around the equator of the cell during interphase.
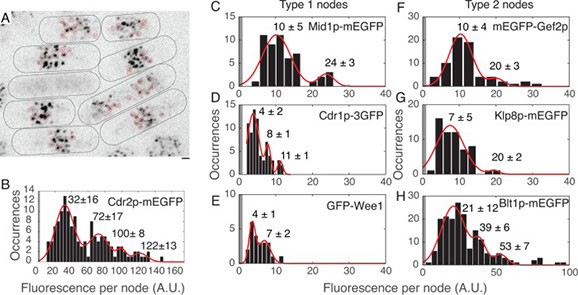
Full text
2016
C. Laplante, F. Huang, I.R. Tebbs, J. Bewersdorf, T.D. Pollard
“Molecular organization of cytokinesis nodes and contractile rings by super-resolution fluorescence microscopy of live fission yeast”, PNAS 113(40):E5876-E5885. (2016)
Cytokinesis in animals, fungi, and amoebas depends on the constriction of a contractile ring built from a common set of conserved proteins. Many fundamental questions remain about how these proteins organize to generate the necessary tension for cytokinesis. Using quantitative high-speed fluorescence photoactivation localization microscopy (FPALM), we probed this question in live fission yeast cells at unprecedented resolution. We show that nodes, protein assembly precursors to the contractile ring, are discrete structural units with stoichiometric ratios and distinct distributions of constituent proteins. Anillin Mid1p, Fes/CIP4 homology-Bin/amphiphysin/Rvs (F-BAR) Cdc15p, IQ motif containing GTPase-activating protein (IQGAP) Rng2p, and formin Cdc12p form the base of the node that anchors the ends of myosin II tails to the plasma membrane, with myosin II heads extending into the cytoplasm. This general node organization persists in the contractile ring where nodes move bidirectionally during constriction. We observed the dynamics of the actin network during cytokinesis, starting with the extension of short actin strands from nodes, which sometimes connected neighboring nodes. Later in cytokinesis, a broad network of thick bundles coalesced into a tight ring around the equator of the cell. The actin ring was ∼125 nm wide and ∼125 nm thick. These observations establish the organization of the proteins in the functional units of a cytokinetic contractile ring.
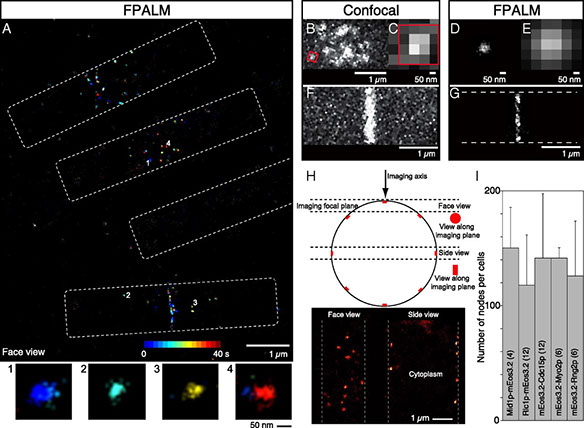
Full text
Fang Huang*, George Sirinakis*, Edward S. Allgeyer, Lena K. Schroeder, Whitney C. Duim, Emil B. Kromann, Thomy Phan, Felix E. Rivera-Molina, Jordan R. Myers, Irnov Irnov, Mark Lessard, Yongdeng Zhang, Mary Ann Handel, Christine Jacobs-Wagner, C. Patrick Lusk, James E. Rothman, Derek Toomre, Martin J. Booth, Joerg Bewersdorf
Ultra-High Resolution 3D Imaging of Whole Cells
2016, Cell 166, 1-13 (2016) *equal authorship
Fluorescence nanoscopy, or super-resolution microscopy, has become an important tool in cell biological research. However, because of its usually inferior resolution in the depth direction (50–80 nm) and rapidly deteriorating resolution in thick samples, its practical biological application has been effectively limited to two dimensions and thin samples. Here, we present the development of whole-cell 4Pi single-molecule switching nanoscopy (W-4PiSMSN), an optical nanoscope that allows imaging of three-dimensional (3D) structures at 10- to 20-nm resolution throughout entire mammalian cells. We demonstrate the wide applicability of W-4PiSMSN across diverse research fields by imaging complex molecular architectures ranging from bacteriophages to nuclear pores, cilia, and synaptonemal complexes in large 3D cellular volumes.
Full text
Florencia del Viso*, Fang Huang*, Jordan Myers, Madeleine Chalfant, Yongdeng Zhang, Nooreen Reza, Joerg Bewersdorf, C. Patrick Lusk, Mustafa K. Khokha
Congenital Heart Disease Genetics Uncovers Context-Dependent Organization and Function of Nucleoporins at Cilia
Developmental Cell, 38, 478-492 (2016) * equal authorship
Human genomics is identifying candidate genes for congenital heart disease (CHD), but discovering the underlying mechanisms remains challenging. In a patient with CHD and heterotaxy (Htx), a disorder of left-right patterning, we previously identified a duplication in Nup188. However, a mechanism to explain how a component of the nuclear pore complex (NPC) could cause Htx/CHD was undefined. Here, we show that knockdown of Nup188 or its binding partner Nup93 leads to a loss of cilia during embryonic development while leaving NPC function largely intact. Many data, including the localization of endogenous Nup188/93 at cilia bases, support their direct role at cilia. Super-resolution imaging of Nup188 shows two barrel-like structures with dimensions and organization incompatible with an NPC-like ring, arguing against a proposed “ciliary pore complex.” We suggest that the nanoscale organization and function of nucleoporins are context dependent in a way that is required for the structure of the heart.
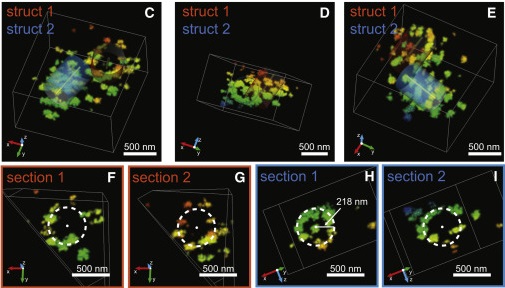
Full text
Francesca Bottanelli*, Emil B. Kromann*, Edward S. Allgeyer, Roman S. Erdmann, Stephanie Wood Baguley, George Sirinakis, Alanna Schepartz, David Baddeley, Derek K. Toomre, James E. Rothman & Joerg Bewersdorf
Two-colour live-cell nanoscale imaging of intracellular targets
Nature Communications, 7, Article number:10778 (2016) * equal authorship
Stimulated emission depletion (STED) nanoscopy allows observations of subcellular dynamics at the nanoscale. Applications have, however, been severely limited by the lack of a versatile STED-compatible two-colour labelling strategy for intracellular targets in living cells. Here we demonstrate a universal labelling method based on the organic, membrane-permeable dyes SiR and ATTO590 as Halo and SNAP substrates. SiR and ATTO590 constitute the first suitable dye pair for two-colour STED imaging in living cells below 50 nm resolution. We show applications with mitochondria, endoplasmic reticulum, plasma membrane and Golgi-localized proteins, and demonstrate continuous acquisition for up to 3 min at 2-s time resolution.
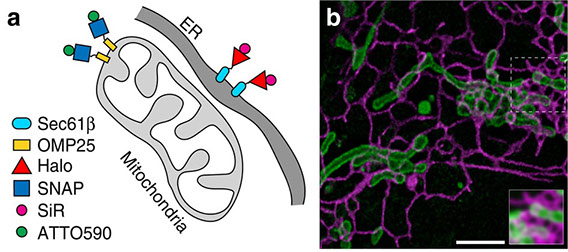
Full text
Jacopo Antonello, Emil B Kromann, Daniel Burke, Joerg Bewersdorf, Martin J Booth
Coma aberrations in combined two-and three-dimensional STED nanoscopy
Optics Letters 41 (15), 3631-3634 (2016)
Stimulated emission depletion (STED) microscopes, like all super-resolution methods, are sensitive to aberrations. Of particular importance are aberrations that affect the quality of the depletion focus, which requires a point of near-zero intensity surrounded by strong illumination. We present analysis, modeling, and experimental measurements that show the effects of coma aberrations on depletion patterns of two-dimensional (2D) and three-dimensional (3D) STED configurations. Specifically, we find that identical coma aberrations create focal shifts in opposite directions in 2D and 3D STED. This phenomenon could affect the precision of microscopic measurements and has ramifications for the efficacy of combined 2D/3D STED systems.
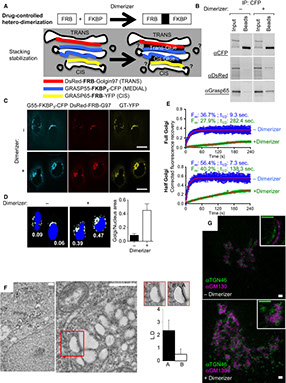
Julia Dancourt, Hong Zheng, Francesca Bottanelli, Edward S Allgeyer, Joerg Bewersdorf, Morven Graham, Xinran Liu, James E Rothman, Grégory Lavieu
Small cargoes pass through synthetically glued Golgi stacks
FEBS letters Vol. 590, Issue 12, 1675-1686 (2016)
How are proteins transported across the stacked cisternae of the Golgi apparatus? Do they stay within the cisterna while the latter matures and progresses in an anterograde manner, or do they navigate between the cisternae via vesicles? Using synthetic biology, we engineered new tools designed to stabilize intercisternal adhesion such that Golgi cisternae are literally glued together, thus preventing any possible cisternal progression. Using bulk secretory assays and single-cell live imaging, we observed that small cargoes (but not large aggregated cargoes including collagen) still transited through glued Golgi, although the rate of transport was moderately reduced. ARF1, whose membrane recruitment is required for budding COPI vesicles, continues to cycle on and off glued Golgi. Numerous COPI-size vesicles were intercalated among the glued Golgi cisternae. These results suggest that cisternal progression is not required for anterograde transport, but do not address the possibility of cisternal maturation in situ.
Full text
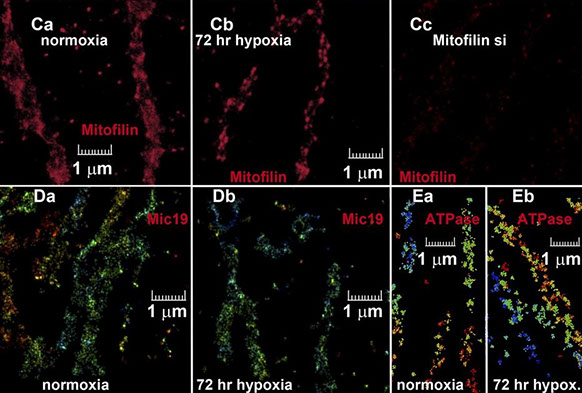
Lydie Plecitá-Hlavatá, Hana Engstová, Lukáš Alán, Tomáš Špaček, Andrea Dlasková, Katarína Smolková, Jitka Špačková, Jan Tauber, Vendula Strádalová, Jan Malínský, Mark Lessard, Joerg Bewersdorf, Petr Ježek
Hypoxic HepG2 cell adaptation decreases ATP synthase dimers and ATP production in inflated cristae by mitofilin down-regulation concomitant to MICOS clustering
The FASEB Journal 30 (5), 1941-1957 (2016)
The relationship of the inner mitochondrial membrane (IMM) cristae structure and intracristal space (ICS) to oxidative phosphorylation (oxphos) is not well understood. Mitofilin (subunit Mic60) of the mitochondrial contact site and cristae organizing system (MICOS) IMM complex is attached to the outer membrane (OMM) via the sorting and assembly machinery/topogenesis of mitochondrial outer membrane β-barrel proteins (SAM/TOB) complex and controls the shape of the cristae. ATP synthase dimers determine sharp cristae edges, whereas trimeric OPA1 tightens ICS outlets. Metabolism is altered during hypoxia, and we therefore studied cristae morphology in HepG2 cells adapted to 5% oxygen for 72 h. Three dimensional (3D), super-resolution biplane fluorescence photoactivation localization microscopy with Eos-conjugated, ICS-located lactamase-β indicated hypoxic ICS expansion with an unchanged OMM (visualized by Eos-mitochondrial fission protein-1). 3D direct stochastic optical reconstruction microscopy immunocytochemistry revealed foci of clustered mitofilin (but not MICOS subunit Mic19) in contrast to its even normoxic distribution. Mitofilin mRNA and protein decreased by ∼20%. ATP synthase dimers vs. monomers and state-3/state-4 respiration ratios were lower during hypoxia. Electron microscopy confirmed ICS expansion (maximum in glycolytic cells), which was absent in reduced or OMM-detached cristae of OPA1- and mitofilin-silenced cells, respectively. Hypoxic adaptation is reported as rounding sharp cristae edges and expanding cristae width (ICS) by partial mitofilin/Mic60 down-regulation. Mitofilin-depleted MICOS detaches from SAM while remaining MICOS with mitofilin redistributes toward higher interdistances. This phenomenon causes partial oxphos dormancy in glycolytic cells via disruption of ATP synthase dimers.—Plecitá-Hlavatá, L., Engstová, H., Alán, L., Špaček, T., Dlasková, A., Smolková, K., Špačková, J., Tauber, J., Strádalová, V., Malínský, J., Lessard, M., Bewersdorf, J., Ježek, P. Hypoxic HepG2 cell adaptation decreases ATP synthase dimers and ATP production in inflated cristae by mitofilin down-regulation concomitant to MICOS clustering.
Full text
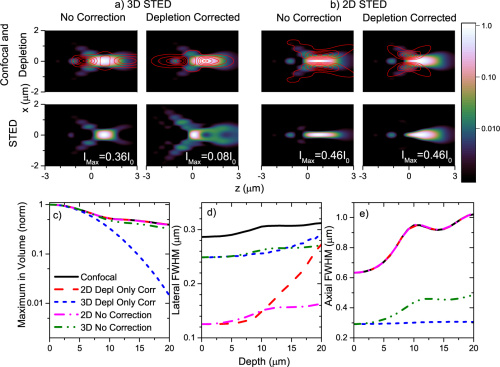
BR Patton, D Burke, D Owald, TJ Gould, J Bewersdorf, MJ Booth
Three-dimensional STED microscopy of aberrating tissue using dual adaptive optics
Optics Express 24 (8), 8862-8876 (2016)
When imaging through tissue, the optical inhomogeneities of the sample generate aberrations that can prevent effective Stimulated Emission Depletion (STED) imaging. This is particularly problematic for 3D-enhanced STED. We present here an adaptive optics implementation that incorporates two adaptive optic elements to enable correction in all beam paths, allowing performance improvement in thick tissue samples. We use this to demonstrate 3D STED imaging of complex structures in Drosophila melanogaster brains.
Full text
C. Laplante, F. Huang, J. Bewersdorf, T. D. Pollard
High-speed super-resolution imaging of live fission yeast cells
Methods in Molecular Biology, Vol. 1390, pp. 45-57 (2016)
We describe a step-by-step method for high-speed fluorescence photoactivation localization microscopy (FPALM) of live fission yeast cells. The resolution with this method is tenfold better than spinning disk confocal microscopy.

Full text
2015
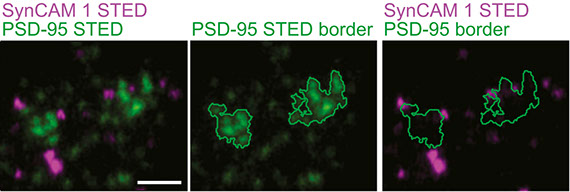
K. P. de Arce, N. Schrod, S. W. R. Metzbower, E. Allgeyer, G. K. ‐W. Kong, A. Tang,
A. J. Krupp, V. Stein, X. Liu, J. Bewersdorf, T. A. Blanpied, V. Lucic, T. Biederer
Topographic mapping of the synaptic cleft into adhesive nanodomains
Neuron, Vol. 88, No. 6, pp. 1165-1172 (2015)
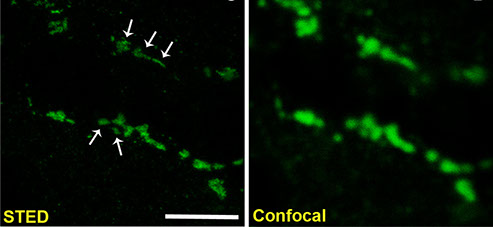
The cleft is an integral part of synapses, yet its macromolecular organization remains unclear. We show here that the cleft of excitatory synapses exhibits a distinct density profile as measured by cryoelectron tomography (cryo-ET). Aiming for molecular insights, we analyzed the synapse-organizing proteins Synaptic Cell Adhesion Molecule 1 (SynCAM 1) and EphB2. Cryo-ET of SynCAM 1 knockout and overexpressor synapses showed that this immunoglobulin protein shapes the cleft’s edge. SynCAM 1 delineates the postsynaptic perimeter as determined by immunoelectron microscopy and super-resolution imaging. In contrast, the EphB2 receptor tyrosine kinase is enriched deeper within the postsynaptic area. Unexpectedly, SynCAM 1 can form ensembles proximal to postsynaptic densities, and synapses containing these ensembles were larger. Postsynaptic SynCAM 1 surface puncta were not static but became enlarged after a long-term depression paradigm. These results support that the synaptic cleft is organized on a nanoscale into sub-compartments marked by distinct trans-synaptic complexes.
Full text
M. G. M. Velasco, E. S. Allgeyer, P. Yuan, J. Grutzendler, J. Bewersdorf
Absolute two-photon excitation spectra of red and far-red fluorescent probes
Optics Letters, Vol. 40, No. 21, pp. 4915-4918 (2015)
Efficient use of two-photon excitation (TPE) microscopy requires knowledge of the absolute TPE action cross sections (ATACSs) of fluorescent probes. However, these values are not available for recently developed dyes, which exhibit superior properties in many modern microscopy applications. We report ATACSs of five red to far-red organic dyes, ATTO 647N, STAR 635P, silicon rhodamine, ATTO 594, and ATTO 590. The dyes were found to have large ATACSs (>100 GM) at their respective wavelength peaks, thus supporting their use as bright fluorescent markers in TPE microscopy.
Full text
X. Hao, E. S. Allgeyer, M. J. Booth, J. Bewersdorf
Point-spread function optimization in isoSTED nanoscopy
Optics Letters, Vol. 40, No. 15, pp. 3627-3630 (2015)
IsoSTED nanoscopy, a variant of stimulated emission depletion (STED) microscopy, utilizes two opposing objective lenses and features the highest three-dimensional resolution of STED nanoscopes currently available. However, this technique is limited by axially repetitive side minima in the interference pattern of the depletion point-spread function (PSF), which can lead to ghost images. Here, we describe novel strategies to further improve the performance of isoSTED nanoscopy by reshaping the PSF. In particular, we propose employing moderate defocus on the depletion beam to reduce the side minima. Furthermore, we demonstrate a simplified alternative based on objective misalignment and quantitatively compare the expected performance between the two approaches.
Full text

Y. Lin, J. J. Long, F. Huang, W.C. Duim, S. Kirschbaum, Y. Zhang, L. K. Schroeder, A. A. Rebane,
M. G. Velasco, A. Virrueta, D. W. Moonan, J. Jiao, S. Y. Hernandez, Y. Zhang, J. Bewersdorf
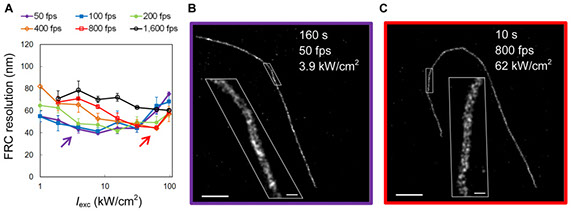
Quantifying and optimizing single-molecule switching nanoscopy at high speeds
PLOS ONE, DOI: 10(5):e0128135 (2015)
Single-molecule switching nanoscopy overcomes the diffraction limit of light by stochastically switching single fluorescent molecules on and off, and then localizing their positions individually. Recent advances in this technique have greatly accelerated the data acquisition speed and improved the temporal resolution of super-resolution imaging. However, it has not been quantified whether this speed increase comes at the cost of compromised image quality. The spatial and temporal resolution depends on many factors, among which laser intensity and camera speed are the two most critical parameters. Here we quantitatively compare the image quality achieved when imaging Alexa Fluor 647-immunolabeled microtubules over an extended range of laser intensities and camera speeds using three criteria – localization precision, density of localized molecules, and resolution of reconstructed images based on Fourier Ring Correlation. We found that, with optimized parameters, single-molecule switching nanoscopy at high speeds can achieve the same image quality as imaging at conventional speeds in a 5–25 times shorter time period. Furthermore, we measured the photoswitching kinetics of Alexa Fluor 647 from single-molecule experiments, and, based on this kinetic data, we developed algorithms to simulate single-molecule switching nanoscopy images. We used this software tool to demonstrate how laser intensity and camera speed affect the density of active fluorophores and influence the achievable resolution. Our study provides guidelines for choosing appropriate laser intensities for imaging Alexa Fluor 647 at different speeds and a quantification protocol for future evaluations of other probes and imaging parameters.
Full text
D. Burke, B. Patton, F. Huang, J. Bewersdorf, M. J. Booth
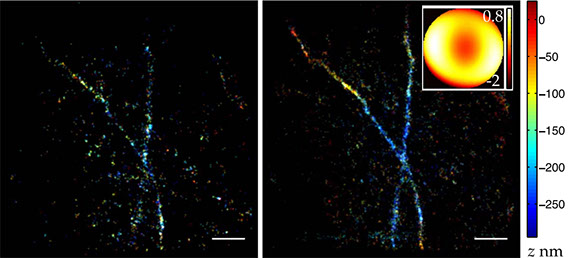
Adaptive optics correction of specimen-induced aberrations in single-molecule switching microscopy
Optica, Vol. 2, No. 2, pp. 177-185 (2015)
Single-molecule switching (SMS) microscopy is a super-resolution method capable of producing images with resolutions far exceeding that of the classical diffraction limit. However, like all optical microscopes, SMS microscopes are sensitive to, and often limited by, specimen-induced aberrations. Adaptive optics (AO) has proven beneficial in a range of microscopes to overcome the limitations caused by aberrations. We report here on new AO methods for SMS microscopy that enable the feedback correction of specimeninduced aberrations. The benefits are demonstrated through two-dimensional and three-dimensional STORM imaging. We expect that this advance will broaden the scope of SMS microscopy by enabling deep-cell and tissue-level imaging.
Full text
2014
M. J. Amaya, A. G. Oliveira, L. K. Schroeder, E. S. Allgeyer, J. Bewersdorf, M. H. Nathanson
Apical localization of inositol 1,4,5-trisphosphate receptors
is independent of extended synaptotagmins in hepatocytes
PLoS One, DOI: 10.1371/journal.pone.0114043 (2014)
Extended synaptotagmins (E-Syts) are a recently identified family of proteins that tether the endoplasmic reticulum (ER) to the plasma membrane (PM) in part by conferring regulation of cytosolic calcium (Ca2+) at these contact sites (Cell, 2013). However, the mechanism by which E-Syts link this tethering to Ca2+ signaling is unknown. Ca2+ waves in polarized epithelia are initiated by inositol 1,4,5-trisphosphate receptors (InsP3Rs), and these waves begin in the apical region because InsP3Rs are targeted to the ER adjacent to the apical membrane. In this study we investigated whether E-Syts are responsible for this targeting. Primary rat hepatocytes were used as a model system, because a single InsP3R isoform (InsP3R-II) is tethered to the peri-apical ER in these cells. Additionally, it has been established in hepatocytes that the apical localization of InsP3Rs is responsible for Ca2+ waves and secretion and is disrupted in disease states in which secretion is impaired. We found that rat hepatocytes express two of the three identified E-Syts (E-Syt1 and E-Syt2). Individual or simultaneous siRNA knockdown of these proteins did not alter InsP3R-II expression levels, apical localization or average InsP3R-II cluster size. Moreover, apical secretion of the organic anion 5-chloromethylfluorescein diacetate (CMFDA) was not changed in cells lacking E-Syts but was reduced in cells in which cytosolic Ca2+ was buffered. These data provide evidence that E-Syts do not participate in the targeting of InsP3Rs to the apical region. Identifying tethers that bring InsP3Rs to the apical region remains an important question, since mis-targeting of InsP3Rs leads to impaired secretory activity.
Full text
P. D. Uchil, T. Pawliczek, T. D. Reynolds, S. Ding, A. Hinz, J. B. Munro, F. Huang, R. W. Floyd,
H. Yang, W. L. Hamilton, J. Bewersdorf, Y. Xiong, D. A. Calderwood, W. Mothes
TRIM15 is a focal adhesion protein that regulates focal adhesion disassembly
Journal of Cell Science, DOI: 10.1242/jcs.143537 (2014)
Focal adhesions (FAs) are macromolecular complexes that connect the actin cytoskeleton to the extracellular matrix. Dynamic turnover of FAs is critical for cell migration. Paxillin is a multi-adaptor protein that plays an important role in regulating FA dynamics. Here, we identify TRIM15, a member of the TRIpartite Motif protein family, as a paxillin-interacting factor and a component of FAs. TRIM15 localizes to focal contacts in a myosin II-independent manner by an interaction between its coiled coil domain and the LD2 motif of paxillin. Unlike other FA proteins, TRIM15 is a stable FA component with restricted mobility due to its ability to form oligomers. TRIM15-depleted cells display impaired cell migration and FA disassembly rates in addition to enlarged FAs. Thus, our studies demonstrate a cellular function for TRIM15 as a regulatory component of FA turnover and cell migration.
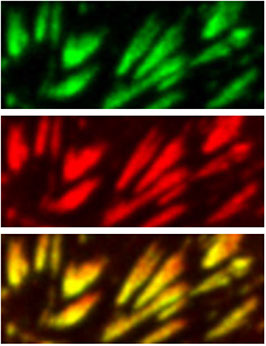
Full text
J. H. Henson, A. D. Gianakas, L. H. Henson, C. L. Lakin, M. K. Voss, J. Bewersdorf, R. Oldenbourg, R. L. Morris
Broadening the spectrum of actin-based protrusive activity mediated by Arp2/3 complex-facilitated polymerization:
Motility of cytoplasmic ridges and tubular projections
Cytoskeleton, Vol. 71, pp. 484–500 (2014)
Arp2/3 complex-facilitated actin polymerization plays an essential role in a variety of cellular functions including motility, adherence, endocytosis, and trafficking. In the present study, we employ the sea urchin coelomocyte experimental model system to test the hypotheses that Arp2/3 complex-nucleated actin assembly mediates the motility of two unusual cellular protrusions; the cytoplasmic ridges present during coelomocyte spreading, and inducible, tubular-shaped, and neurite-like projections. Our investigations couple pharmacological manipulation employing inhibitors of actin polymerization and the Arp2/3 complex with a wide array of imaging methods including digitally enhanced phase contrast, DIC, and polarization light microscopy of live cells; conventional, confocal and super-resolution light microscopy of fluorescently labeled cells; and scanning and transmission electron microscopy. Taken together, the results of this study indicate that Arp2/3 complexfacilitated actin polymerization underlies the mo lity of coelomocyte cytoplasmic ridges and tubular projections, that these processes are related to each other, and that they have been preliminarily identified in other cell types. The results also highlight the broad spectrum of actin-based protrusive activities dependent on the Arp2/ 3 complex and provide additional insights into the pervasive nature of this ubiquitous actin nucleator. Furthermore, we provide the first evidence of a possible mechanistic difference between the impacts of the small molecule drugs BDM and CK666 on the Arp2/3 complex.
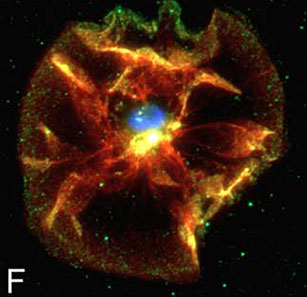
Full text
R. S. Erdmann, H. Takakura, A. D. Thompson, F. Rivera-Molina,
E. S. Allgeyer, J. Bewersdorf, D. Toomre D, A. SchepartzSuper-resolution imaging of the Golgi in live cells with a bioorthogonal ceramide probe
Angewandte Chemie International Edition, DOI: 10.1002/anie.201403349 (2014)
We report a lipid-based strategy to visualize Golgi structure and dynamics at super-resolution in live cells. The method is based on two novel reagents: a trans-cyclooctene-containing ceramide lipid (Cer-TCO) and a highly reactive, tetrazine-tagged near-IR dye (SiR-Tz). These reagents assemble via an extremely rapid "tetrazine-click" reaction into Cer-SiR, a highly photostable "vital dye" that enables prolonged live-cell imaging of the Golgi apparatus by 3D confocal and STED microscopy. Cer-SiR is nontoxic at concentrations as high as 2 μM and does not perturb the mobility of Golgi-resident enzymes or the traffic of cargo from the endoplasmic reticulum through the Golgi and to the plasma membrane.
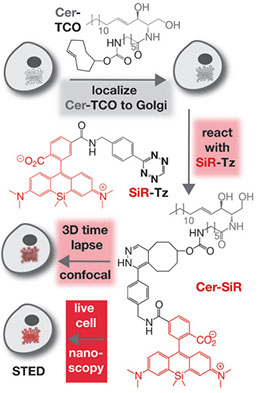
Full text
M. Stagi, Z. A. Klein, T. J. Gould, J. Bewersdorf, S. M. Strittmatter
Lysosome size, motility and stress response regulated by fronto-temporal dementia modifier TMEM106B
Molecular and Cellular Neuroscience, DOI: 10.1016/j.mcn.2014.07.006 (2014)
Fronto-temporal lobar degeneration with TDP-43 (FTLD-TDP) is a fatal neurodegeneration. TMEM106B variants are linked to FTLD-TDP risk, and TMEM106B is lysosomal. Here, we focus on neuronal TMEM106B, and demonstrate co-localization and traffic with lysosomal LAMP-1. pH-sensitive reporters demonstrate that the TMEM106B C-terminus is lumenal. The TMEM106B N-terminus interacts with endosomal adaptors and other TMEM106 proteins. TMEM106B knockdown reduces neuronal lysosomal number and diameter by STED microscopy, and overexpression enlarges LAMP-positive structures. Reduction of TMEM106B increases axonally transported lysosomes, while TMEM106B elevation inhibits transport and yields large lysosomes in the soma. TMEM106B overexpression alters lysosomal stress signaling, causing a translocation of the mTOR-sensitive transcription factor, TFEB, to neuronal nuclei. TMEM106B loss-of-function delays TFEB translocation after Torin-1-induced stress. Enlarged TMEM106B-overexpressing lysosomes maintain organelle integrity longer after lysosomal photodamage than do control lysosomes, while small TMEM106B-knockdown lysosomes are more sensitive to illumination. Thus, neuronal TMEM106B plays a central role in regulating lysosomal size, motility and responsiveness to stress, highlighting the possible role of lysosomal biology in FTLD-TDP.
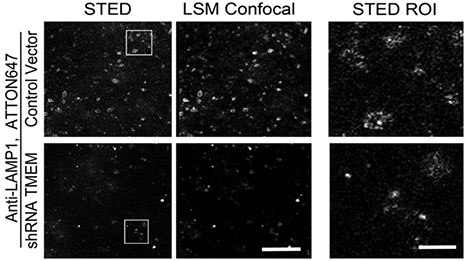
Full text
H. C. Lim, I. V. Surovtsev, B. G. Beltran, F. Huang, J. Bewersdorf, C. Jacobs-Wagner
Evidence for a DNA-relay mechanism in ParABS-mediated chromosome segregation
eLife, 2014;10.7554/eLife.02758 (2014)
The widely conserved ParABS system plays a major role in bacterial chromosome segregation. How the components of this system work together to generate translocation force and directional motion remains uncertain. Here, we combine biochemical approaches, quantitative imaging and mathematical modeling to examine the mechanism by which ParA drives the translocation of the ParB/parS partition complex in Caulobacter crescentus. Our experiments, together with simulations grounded on experimentally-determined biochemical and cellular parameters, suggest a novel 'DNA-relay' mechanism in which the chromosome plays a mechanical function. In this model, DNA-bound ParA-ATP dimers serve as transient tethers that harness the elastic dynamics of the chromosome to relay the partition complex from one DNA region to another across a ParA-ATP dimer gradient. Since ParA-like proteins are implicated in the partitioning of various cytoplasmic cargos, the conservation of their DNA-binding activity suggests that the DNA-relay mechanism may be a general form of intracellular transport in bacteria.
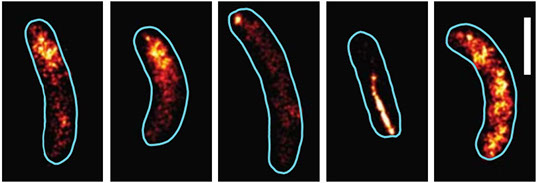
Full text
S. Nath, J. Dancourt, V. Shteyn, G. Puente, W. M. Fong, S. Nag,
J. Bewersdorf, A. Yamamoto, B. Antonny, T. J. Melia
Lipidation of the LC3/GABARAP family of autophagy proteins relies on a membrane-curvature-sensing domain in Atg3
Nature Cell Biology, Vol. 16, pp. 415–424 (2014)
The components supporting autophagosome growth on the cup-like isolation membrane are likely to be different from those found on closed and maturing autophagosomes. The highly curved rim of the cup may serve as a functionally required surface for transiently associated components of the early acting autophagic machinery. Here we demonstrate that the E2-like enzyme, Atg3, facilitates LC3/GABARAP lipidation only on membranes exhibiting local lipid-packing defects. This activity requires an amino-terminal amphipathic helix similar to motifs found on proteins targeting highly curved intracellular membranes. By tuning the hydrophobicity of this motif, we can promote or inhibit lipidation in vitro and in rescue experiments in Atg3-knockout cells, implying a physiologic role for this stress detection. The need for extensive lipid-packing defects suggests that Atg3 is designed to work at highly curved membranes, perhaps including the limiting edge of the growing phagophore.
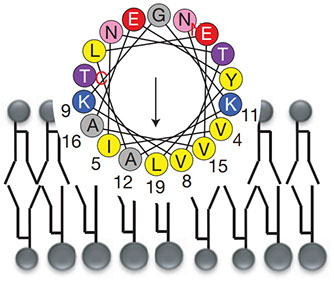
Full text
H. Deschout, F. Cella Zanacchi, A. Diaspro, J. Bewersdorf, S. T. Hess, K. Braeckmans
Precisely and accurately localizing single emitters in fluorescence microscopy
Nature Methods, Vol. 11, No. 3, pp. 253-266 (2014)
Methods based on single-molecule localization and photophysics have brought nanoscale imaging with visible light into reach. This has enabled single-particle tracking applications for studying the dynamics of molecules and nanoparticles and contributed to the recent revolution in super-resolution localization microscopy techniques. Crucial to the optimization of such methods are the precision and accuracy with which single fluorophores and nanoparticles can be localized. We present a lucid synthesis of the developments on this localization precision and accuracy and their practical implications in order to guide the increasing number of researchers using single-particle tracking and super-resolution localization microscopy.
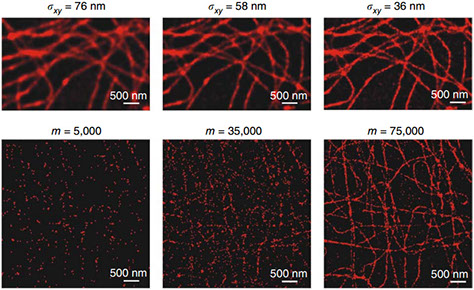
Full text
F. Wilfling, A. R. Thiam, M. Olarte, J. Wang, R. Beck, T. J. Gould, E. S. Allgeyer,
F. Pincet, J. Bewersdorf, R. V. Farese Jr, T. C. Walther
Arf1/COPI machinery acts directly on lipid droplets and enables their connection to the ER for protein targeting
eLife, eLife 2014;3:e01607 (2014)
Lipid droplets (LDs) are ubiquitous organelles that store neutral lipids, such as triacylglycerol (TG), as reservoirs of metabolic energy and membrane precursors. The Arf1/COPI protein machinery, known for its role in vesicle trafficking, regulates LD morphology, targeting of specific proteins to LDs and lipolysis through unclear mechanisms. Recent evidence shows that Arf1/COPI can bud nano-LDs (∼60 nm diameter) from phospholipid-covered oil/water interfaces in vitro. We show that Arf1/COPI proteins localize to cellular LDs, are sufficient to bud nano-LDs from cellular LDs, and are required for targeting specific TG-synthesis enzymes to LD surfaces. Cells lacking Arf1/COPI function have increased amounts of phospholipids on LDs, resulting in decreased LD surface tension and impairment to form bridges to the ER. Our findings uncover a function for Arf1/COPI proteins at LDs and suggest a model in which Arf1/COPI machinery acts to control ER-LD connections for localization of key enzymes of TG storage and catabolism.
Full text
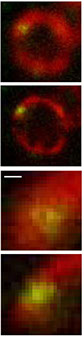
2013
S. Liu, E. B. Kromann, W. D. Krueger, J. Bewersdorf, K. A. Lidke
Three dimensional single molecule localization using a phase retrieved pupil function
Optics Express, 21:29462-29487 (2013)
Localization-based superresolution imaging is dependent on finding the positions of individual fluorophores in a sample by fitting the observed single-molecule intensity pattern to the microscope point spread function (PSF). For three-dimensional imaging, system-specific aberrations of the optical system can lead to inaccurate localizations when the PSF model does not account for these aberrations. Here we describe the use of phase-retrieved pupil functions to generate a more accurate PSF and therefore more accurate 3D localizations. The complex-valued pupil function contains information about the system-specific aberrations and can thus be used to generate the PSF for arbitrary defocus. Further, it can be modified to include depth dependent aberrations. We describe the phase retrieval process, the method for including depth dependent aberrations, and a fast fitting algorithm using graphics processing units. The superior localization accuracy of the pupil function generated PSF is demonstrated with dual focal plane 3D superresolution imaging of biological structures.
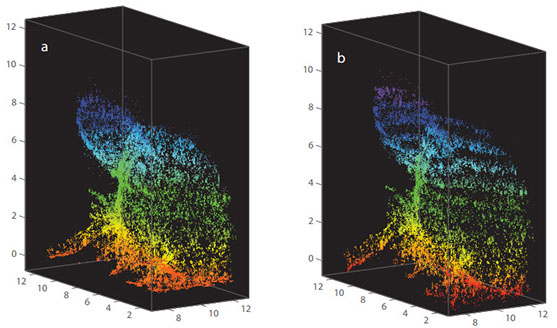
Full text
P. A. Pellett, F. Dietrich, J. Bewersdorf, J. E. Rothman, G. Lavieu
Inter-Golgi transport mediated by COPI-containing vesicles carrying small cargoes
eLife, 2013;2:e01296 (2013)
A core prediction of the vesicular transport model is that COPI vesicles are responsible for trafficking anterograde cargoes forward. In this study, we test this prediction by examining the properties and requirements of inter-Golgi transport within fused cells, which requires mobile carriers in order for exchange of constituents to occur. We report that both small soluble and membrane-bound secretory cargo and exogenous Golgi resident glycosyl-transferases are exchanged between separated Golgi. Large soluble aggregates, which traverse individual stacks, do not transfer between Golgi, implying that small cargoes (which can fit in a typical transport vesicle) are transported by a different mechanism. Super-resolution microscopy reveals that the carriers of both anterograde and retrograde cargoes are the size of COPI vesicles, contain coatomer, and functionally require ARF1 and coatomer for transport. The data suggest that COPI vesicles traffic both small secretory cargo and steady-state Golgi resident enzymes among stacked cisternae that are stationary.
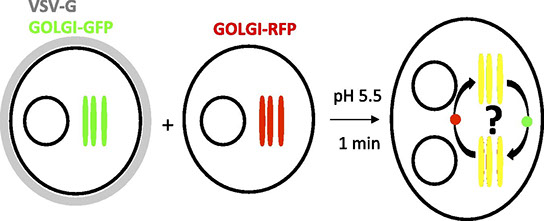
Full text
T. M. P. Hartwich, F. V. Subach, L. Cooley, V. V. Verkhusha, J. Bewersdorf,
Determination of two-photon photoactivation rates of fluorescent proteins
Physical Chemistry Chemical Physics, DOI: 10.1039/C3CP51035B (2013)
The application of two-photon activation of photoactivatable fluorescent proteins is limited by a lack of information about two-photon activation rates. Here we present rates for the commonly used photoactivatable proteins PAmCherry, PAmKate and PA-GFP at different wavelengths using a novel method that allows us to determine the two-photon activation rates directly, independent of any reference data, with microscopic sample volumes. We show that PAmCherry features the highest rates of the tested proteins at 700 nm activation wavelength followed by PAmKate. Towards longer wavelengths, two-photon activation rates decrease for all three proteins. For PAmCherry, our data contradicts an activation model relying solely on two-photon activation and suggests additional activation pathways requiring at least two absorption steps. Our method is readily expandable to other photoactivatable fluorescent molecules. The presented results allow optimization of experimental conditions in spectroscopic and imaging techniques such as super-resolution fluorescence microscopy.
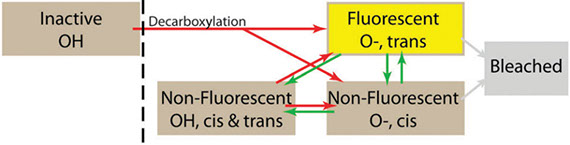
Full text
F. Huang, T. M. P. Hartwich, F. E. Rivera-Molina, Y. Lin, W. C. Duim, J. J. Long, P. D. Uchil, J. R. Myers,
M. A. Baird, W. Mothes, M. W. Davidson, D. Toomre, J. Bewersdorf
Video-Rate Nanoscopy Using sCMOS Camera-Specific Single-Molecule Localization Algorithms
Nature Methods, DOI:10.1038/NMETH.2488 (2013)
Newly developed scientific complementary metal-oxide semiconductor (sCMOS) cameras have the potential to dramatically accelerate data acquisition, enlarge the field of view and increase the effective quantum efficiency in single-molecule switching nanoscopy. However, sCMOS-intrinsic pixel-dependent readout noise substantially lowers the localization precision and introduces localization artifacts. We present algorithms that overcome these limitations and that provide unbiased, precise localization of single molecules at the theoretical limit. Using these in combination with a multi-emitter fitting algorithm, we demonstrate single-molecule localization super-resolution imaging at rates of up to 32 reconstructed images per second in fixed and living cells.
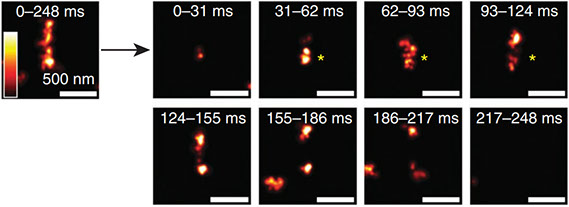
Full text
M. F. Juette, F. E. Rivera-Molina, D. K. Toomre, J. Bewersdorf
Adaptive Optics Enables Three-Dimensional Single Particle Tracking at the Sub-millisecond Scale
Applied Physics Letters, 102:173702 (2013)
We present the integration of an adaptive optics element into a feedback-driven single particle tracking microscope. Our instrument captures three-dimensional (3D) trajectories with down to 130 μs temporal resolution for dynamic studies on the nanoscale. Our 3D beam steering approach tracks particles over an axial range of >6 μm with ∼2 ms mechanical response times and isolates the sample from any tracking motion. Tracking of transport vesicles containing Alexa488-labeled transferrin glycoprotein in living cells demonstrates the speed and sensitivity of our instrument.
Full text
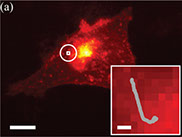
T. J. Gould, E. B. Kromann, D. Burke, M. J. Booth, J. Bewersdorf
An Auto-aligning Stimulated Emission Depletion Microscope Using Adaptive Optics
Optics Letters, 38:1860-1862 (2013)
Stimulated emission depletion (STED) microscopy provides diffraction-unlimited resolution in fluorescence microscopy. Imaging at the nanoscale, however, requires precise alignment of the depletion and excitation laser foci of the STED microscope. We demonstrate here that adaptive optics can be implemented to automatically align STED and confocal images with a precision of 4.3±2.3 nm
Full text
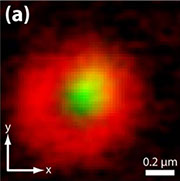
T. J. Gould, P. A. Pellett, J. Bewersdorf
“STED Microscopy” in Fluorescence Microscopy: From Physical Background to Biological Application
Kubitscheck, U. (Ed.), 1st ed., Wiley-VCH Verlag GmbH & Co. KGaA (2013)
A comprehensive introduction to advanced light microscopy methods and their applications. Written in a readily comprehensible style by leading experts and pioneers in the field, this work is designed specifically for students and researchers with no background in physics.
Full text

F. Wilfling, H. Wang, J. T. Haas, N. Krahmer, T. J. Gould, A. Uchida, J. Cheng, M. Graham, R. Christiano,
F. Froehlich, X. Liu, K. K. Buhman, R. A. Coleman, J. Bewersdorf, R. V. Farese, Jr, T. C. Walther
Triacylglycerol Synthesis Enzymes Mediate Lipid Droplet Growth by Relocalizing from the ER to Lipid Droplets
Developmental Cell, 24:1-16, (2013)
Lipid droplets (LDs) store metabolic energy and membrane lipid precursors. With excess metabolic energy, cells synthesize triacylglycerol (TG) and form LDs that grow dramatically. It is unclear how TG synthesis relates to LD formation and growth. Here, we identify two LD subpopulations: smaller LDs of relatively constant size, and LDs that grow larger. The latter population contains isoenzymes for each step of TG synthesis. Glycerol-3-phosphate acyltransferase 4 (GPAT4), which catalyzes the first and rate-limiting step, relocalizes from the endoplasmic reticulum (ER) to a subset of forming LDs, where it becomes stably associated. ER-to-LD targeting of GPAT4 and other LD-localized TG synthesis isozymes is required for LD growth. Key features of GPAT4 ER-to-LD targeting and function in LD growth are conserved between Drosophila and mammalian cells. Our results explain how TG synthesis is coupled with LD growth and identify two distinct LD subpopulations based on their capacity for localized TG synthesis.
Full text
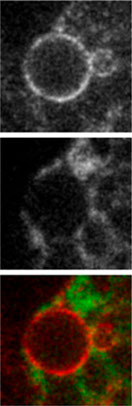
2012
T. J. Gould, J. Bewersdorf
Nanoscopy at Low Light Intensities Shows Its Potential
eLife, 1:e00475 (2012)
A new form of green fluorescent protein allows super-resolution imaging to be performed faster on living cells with low radiation doses.
Full text
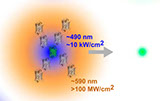
T. J. Gould, D. Burke, J. Bewersdorf, M. J. Booth
Adaptive Optics Enables 3D STED Microscopy in Aberrating Specimens
Optics Express, 20:20998-21009 (2012)
Stimulated emission depletion (STED) microscopy allows fluorescence far-field imaging with diffraction-unlimited resolution. Unfortunately, extending this technique to three-dimensional (3D) imaging of thick specimens has been inhibited by sample-induced aberrations. Here we present the first implementation of adaptive optics in STED microscopy to allow 3D super-resolution imaging in strongly aberrated imaging conditions, such as those introduced by thick biological tissue.
Full text
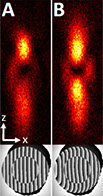

T. J. Gould, S. T. Hess, J. Bewersdorf
Optical Nanoscopy: from Acquisition to Analysis
Annual Review of Biomedical Engineering, 14:231–254 (2012)
The resolution attainable with stimulated emission depletion (STED) microscopy greatly depends on the quality of the STED laser focus. So far, visual inspection of a measured STED focus has been the only convenient means of gauging the source of aberrations. Here we describe a method, requiring no instrument modifications, for obtaining an equivalent to the complex pupil function at the back aperture of the objective and show that it provides quantitative information about aberration sources (including aberrations induced by the objective or sample). We show the accuracy of this field representation to be sufficient for reconstructing the STED focus in three dimensions and determining corrective steps.
Full text
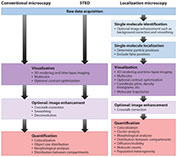
C. Lv, T. J. Gould, J. Bewersdorf, D. Zenisek
High Resolution Optical Imaging of Zebrafish Larval Ribbon Synapse protein
RIBEYE, RIM2, and CaV1.4 by STED Microscopy
Microscopy & Microanalysis, 18:1-8 (2012)
The synaptic ribbon is a unique presynaptic structure with an intricate morphology in photoreceptors. Because of the resolution limit in conventional fluorescence microscopy, investigating ribbon protein locations has been challenging, especially in the early development stages of model animals. Here, we used stimulated emission depletion microscopy, a super-resolution imaging technique, to look at retina sections in 4 days post-fertilization ~dpf! zebrafish. We observed that in photoreceptor cells, RIBEYE and RIM2 are expressed along the synaptic ribbon, with RIM2 consistently located inside of the horseshoe-shaped synaptic ribbon structure with RIBEYE located on the outside. The L-type calcium channel subunit, CACNA1F, exhibited small spot-like staining beneath the RIM2 and RIBEYE structures. Using morpholino antisense oligonucleotides to knock down RIBEYE expression, we observed fewer and shorter ribbons in the photoreceptor outer plexiform layers of 4 dpf fish retina as well as a reduction in RIM2 expression. The clustering of CACNA1F in these blind fish was no longer observed, but instead showed a diffuse expression in the photoreceptor terminal.
Full text
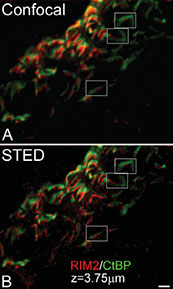
M. Fritsche, L. G. Reinholdt, M. Lessard, M. A. Handel, J. Bewersdorf, D. W. Heermann
The Impact of Entropy on the Spatial Organization of Synaptonemal Complexes within the Cell Nucleus
PLOS One, 7(5):e36282 (2012)
We employ 4Pi-microscopy to study SC organization in mouse spermatocyte nuclei allowing for the three-dimensional reconstruction of the SC's backbone arrangement. Additionally, we model the SCs in the cell nucleus by confined, self-avoiding polymers, whose chain ends are attached to the envelope of the confining cavity and diffuse along it. This work helps to elucidate the role of entropy in shaping pachytene SC organization. The framework provided by the complex interplay between SC polymer rigidity, tethering and confinement is able to qualitatively explain features of SC organization, such as mean squared end-to-end distances, mean squared center-of-mass distances, or SC density distributions. However, it fails in correctly assessing SC entanglement within the nucleus. In fact, our analysis of the 4Pi-microscopy images reveals a higher ordering of SCs within the nuclear volume than what is expected by our numerical model. This suggests that while effects of entropy impact SC organization, the dedicated action of proteins or actin cables is required to fine-tune the spatial ordering of SCs within the cell nucleus.
Full text
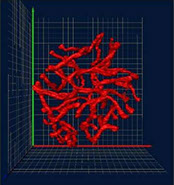
E. B. Kromann, T. J. Gould, M. F. Juette, J. E. Wilhjelm, and J. Bewersdorf
Quantitative Pupil Analysis in STED Microscopy Using Phase Retrieval
Optics Letters, Vol.37, No. 11, pp. 1805-1807 (2012)
The resolution attainable with stimulated emission depletion (STED) microscopy greatly depends on the quality of the STED laser focus. So far, visual inspection of a measured STED focus has been the only convenient means of gauging the source of aberrations. Here we describe a method, requiring no instrument modifications, for obtaining an equivalent to the complex pupil function at the back aperture of the objective and show that it provides quantitative information about aberration sources (including aberrations induced by the objective or sample). We show the accuracy of this field representation to be sufficient for reconstructing the STED focus in three dimensions and determining corrective steps.
Full text
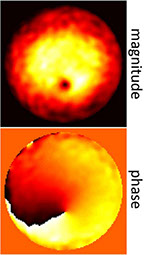
2011
P. A. Pellett, X. Sun, T. J. Gould, J. E. Rothman, M. Xu, I. R. Correa, J. Bewersdorf
Two-Color STED Microscopy in Living Cells
Biomedical Optics Express, 2:2364-2371 (2011)
Diffraction-unlimited resolution provided by Stimulated Emission Depletion (STED) microscopy allows for imaging cellular processes in living cells that are not visible by conventional microscopy. However, it has so far not been possible to study dynamic nanoscale interactions because multicolor live cell STED microscopy has yet to be demonstrated and suitable labeling technologies and protocols are lacking. Here we report the first realization of two-color STED imaging in living cells. Using improved SNAPf and CLIPf technologies to label epidermal growth factor (EGF) and EGF receptor (EGFR), we report resolutions of 78 nm and 82 nm for 22 sequential two-color scans in living cells.
Full text
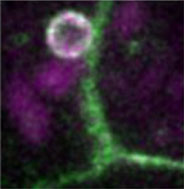
M. J. Mlodzianoski, J. M. Schreiner, S. P. Callahan, K. Smolková, A. Dlasková,
J. Šantorová, P. Ježek, J. Bewersdorf
Sample Drift Correction in 3D Fluorescence Photoactivation Localization Microscopy
Optics Express, 19:15009-15019 (2011)
The recent development of diffraction-unlimited far-field fluorescence microscopy has overcome the classical resolution limit of ~250 nm of conventional light microscopy by about a factor of ten. The improved resolution, however, reveals not only biological structures at an unprecedented resolution, but is also susceptible to sample drift on a much finer scale than previously relevant. Without correction, sample drift leads to smeared images with decreased resolution, and in the worst case to misinterpretation of the imaged structures. This poses a problem especially for techniques such as Fluorescence Photoactivation Localization Microscopy (FPALM/PALM) or Stochastic Optical Reconstruction Microscopy (STORM), which often require minutes recording time. Here we discuss an approach that corrects for three-dimensional (3D) drift in images of fixed samples without the requirement for fiduciary markers or instrument modifications. Drift is determined by calculating the spatial cross-correlation function between subsets of localized particles imaged at different times. Correction down to ~5 nm precision is achieved despite the fact that different molecules are imaged in each frame. We demonstrate the performance of our drift correction algorithm with different simulated structures and analyze its dependence on particle density and localization precision. By imaging mitochondria with Biplane FPALM we show our algorithm’s feasibility in a practical application.
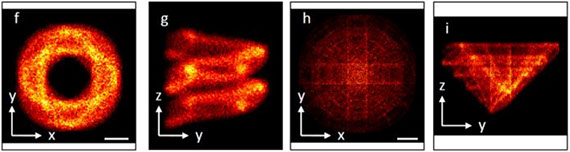

Full text
T. J. Gould, J. R. Myers, J. Bewersdorf
Total Internal Reflection STED Microscopy
Optics Express, 19:13351-1335 (2011)
Diffraction-unlimited resolution provided by Stimulated Emission Depletion (STED) microscopy allows for imaging cellular processes in living cells that are not visible by conventional microscopy. However, it has so far not been possible to study dynamic nanoscale interactions because multicolor live cell STED microscopy has yet to be demonstrated and suitable labeling technologies and protocols are lacking. Here we report the first realization of two-color STED imaging in living cells. Using improved SNAPf and CLIPf technologies to label epidermal growth factor (EGF) and EGF receptor (EGFR), we report resolutions of 78 nm and 82 nm for 22 sequential two-color scans in living cells.
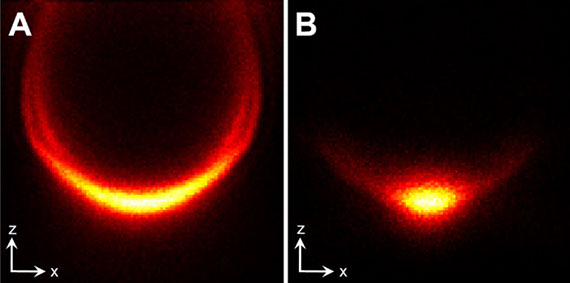

Full text
J. Bewersdorf, R. V. Farese Jr, T. C. Walther
A New Way to Look at Fat
Nature Methods, 8(2):132-133 (2011)
Stimulated Raman scattering (SRS) microscopy is used to directly visualize lipids in cells and model organisms, and facilitates screening for genes involved in fat storage.
Full text
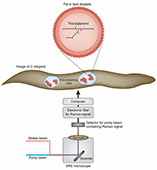
2010
M. F. Juette, J. Bewersdorf
Three-Dimensional Tracking of Single Fluorescent Particles with Submillisecond Temporal Resolution
Nano Letters, 10 (11), 4657–4663 (2010)
Observing dynamics at the nanoscale requires submillisecond time resolution. Notably, in studying biological systems, three-dimensional (3D) trajectories of fluorescently labeled objects such as viruses or transport vesicles often need to be acquired with high temporal resolution. Here, we present a novel instrument that combines scanning-free multiplane detection at 3.2 kHz frame rate and single photon sensitivity with optimized beam-steering capabilities. This setup enables ultrafast 3D localization with submillisecond time resolutionand nanometer localization precision. We demonstrate 3D tracking of single fluorescent particles at speeds of up to 150 nm/ms over several seconds and large volumes. By focused excitation of only the particle of interest, while avoiding confocal pinholes, the system realizes maximum detection efficiency at minimal laser irradiation. These features, combined with the avoidance of stage movement, provide high live-sample compatibility for future biomedical applications.
Full text
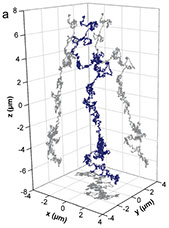
D. Toomre, J. Bewersdorf
A New Wave of Cellular Imaging
Annual Review of Cell and Developmental Biology, 26:285-314 (2010)
Fluorescence imaging methods that push or break the diffraction limit of resolution (approximately 200 nm) have grown explosively. These super-resolution nanoscopy techniques include: stimulated emission depletion (STED), Pointillism microscopy [(fluorescence) photoactivation localization microscopy/stochastic optical reconstruction microscopy, or (F)PALM/STORM], structured illumination, total internal reflection fluorescence microscopy (TIRFM), and those that combine multiple modalities. Each affords unique strengths in lateral and axial resolution, speed, sensitivity, and fluorophore compatibility. We examine the optical principles and design of these new instruments and their ability to see more detail with greater sensitivity—down to single molecules with tens of nanometers resolution. Nanoscopes have revealed transient intermediate states of organelles and molecules in living cells and have led to new discoveries but also biological controversies. We highlight common unifying principles behind nanoscopy such as the conversion of a subset of probes between states (ground or excited) and the use of scanning (ordered or stochastic). We emphasize major advances, biological applications, and promising new developments.
Full text
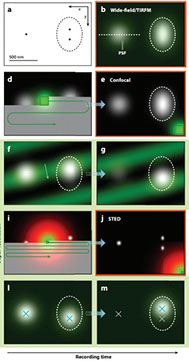
M. F. Juette, T. J. Gould, J. Bewersdorf
“Superresolution Far-Field Fluorescence Microscopy” in Nanobiophotonics
Popescu, G. (Ed.), 1st ed., McGraw-Hill (2010)
Fluorescence imaging methods that push or break the diffraction limit of resolution (approximately 200 nm) have grown explosively. These super-resolution nanoscopy techniques include: stimulated emission depletion (STED), Pointillism microscopy [(fluorescence) photoactivation localization microscopy/stochastic optical reconstruction microscopy, or (F)PALM/STORM], structured illumination, total internal reflection fluorescence microscopy (TIRFM), and those that combine multiple modalities. Each affords unique strengths in lateral and axial resolution, speed, sensitivity, and fluorophore compatibility. We examine the optical principles and design of these new instruments and their ability to see more detail with greater sensitivity—down to single molecules with tens of nanometers resolution. Nanoscopes have revealed transient intermediate states of organelles and molecules in living cells and have led to new discoveries but also biological controversies. We highlight common unifying principles behind nanoscopy such as the conversion of a subset of probes between states (ground or excited) and the use of scanning (ordered or stochastic). We emphasize major advances, biological applications, and promising new developments.
Full text

K. S. Morozova, K. D. Piatkevich, T. J. Gould, J. Zhang, J. Bewersdorf, V. V. Verkhusha
Far-Red Fluorescent Protein Excitable with Red Lasers for Flow Cytometry and Super-Resolution STED Nanoscopy
Biophysical Journal, 99(2):L13-15 (2010)
Far-red fluorescent proteins are required for deep-tissue and whole-animal imaging and multicolor labeling in the red wavelength range, as well as probes excitable with standard red lasers in flow cytometry and fluorescence microscopy. Rapidly evolving superresolution microscopy based on the stimulated emission depletion approach also demands genetically encoded monomeric probes to tag intracellular proteins at the molecular level. Based on the monomeric mKate variant, we have developed a far-red TagRFP657 protein with excitation/emission maxima at 611/657 nm. TagRFP657 has several advantages over existing monomeric far-red proteins including higher photostability, better pH stability, lower residual green fluorescence, and greater efficiency of excitation with red lasers. The red-shifted excitation and emission spectra, as compared to other far-red proteins, allows utilizing TagRFP657 in flow cytometry and fluorescence microscopy simultaneously with orange or near-red fluorescence proteins. TagRFP657 is shown to be an efficient protein tag for the superresolution fluorescence imaging using a commercially available stimulated emission depletion microscope.
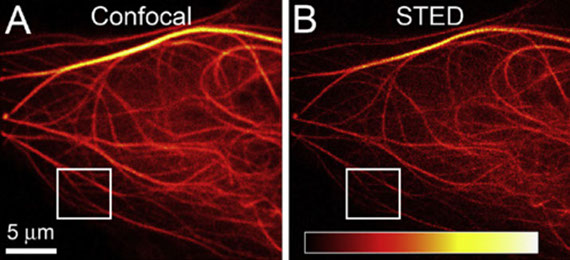
Full text
Some of our earlier work
A. Dlasková, T. Spaček, J. Santorová, L. Plecitá-Hlavatá, Z. Berková, F. Saudek,
M. Lessard, J. Bewersdorf, P. Ježek
4Pi microscopy reveals an impaired three-dimensional mitochondrial network
of pancreatic islet beta-cells, an experimental model of type-2 diabetes
Biochimica et Biophysica Acta - Bioenergetics, 1797:1327-1341 (2010)
Insulin production in pancreatic β-cells is critically linked to mitochondrial oxidative phosphorylation. Increased ATP production triggered by blood glucose represents the β-cells' glucose sensor. Type-2 diabetes mellitus results from insulin resistance in peripheral tissues and impaired insulin secretion. Pathology of diabetic β-cells might be reflected by the altered morphology of mitochondrial network. Its characterization is however hampered by the complexity and density of the three-dimensional (3D) mitochondrial tubular networks in these cell types. Conventional confocal microscopy does not provide sufficient axial resolution to reveal the required details; electron tomography reconstruction of these dense networks is still difficult and time consuming. However, mitochondrial network morphology in fixed cells can also be studied by 4Pi microscopy, a laser scanning microscopy technique which provides an ∼ 7-fold improved axial resolution (∼ 100 nm) over conventional confocal microscopy. Here we present a quantitative study of these networks in insulinoma INS-1E cells and primary β-cells in Langerhans islets. The former were a stably-transfected cell line while the latter were transfected with lentivirus, both expressing mitochondrial matrix targeted redox-sensitive GFP. The mitochondrial networks and their partial disintegration and fragmentation are revealed by carefully created iso-surface plots and their quantitative analysis. We demonstrate that β-cells within the Langerhans islets from diabetic Goto Kakizaki rats exhibited a more disintegrated mitochondrial network compared to those from control Wistar rats and model insulinoma INS-1E cells. Standardization of these patterns may lead to development of morphological diagnostics for Langerhans islets, for the assessment of β-cell condition, before their transplantations.
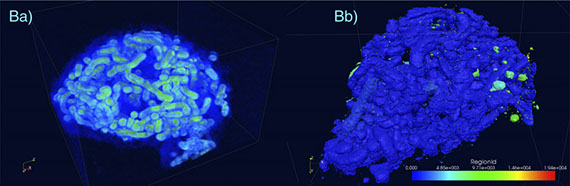
Full text
L. Luo, K. L. Gassman, L. M. Petell, C. L. Wilson, J. Bewersdorf, and L. S. Shopland
The Nuclear Periphery of Embryonic Stem Cells is a Transcriptionally Permissive and Repressive Compartment
Journal of Cell Science, 122(20):3729-3737 (2009)
Fluorescence imaging methods that push or break the diffraction limit of resolution (approximately 200 nm) have grown explosively. These super-resolution nanoscopy techniques include: stimulated emission depletion (STED), Pointillism microscopy [(fluorescence) photoactivation localization microscopy/stochastic optical reconstruction microscopy, or (F)PALM/STORM], structured illumination, total internal reflection fluorescence microscopy (TIRFM), and those that combine multiple modalities. Each affords unique strengths in lateral and axial resolution, speed, sensitivity, and fluorophore compatibility. We examine the optical principles and design of these new instruments and their ability to see more detail with greater sensitivity—down to single molecules with tens of nanometers resolution. Nanoscopes have revealed transient intermediate states of organelles and molecules in living cells and have led to new discoveries but also biological controversies. We highlight common unifying principles behind nanoscopy such as the conversion of a subset of probes between states (ground or excited) and the use of scanning (ordered or stochastic). We emphasize major advances, biological applications, and promising new developments.

Full text
M. J. Mlodzianoski, M. F. Juette, G. L. Beane, J. Bewersdorf
Experimental Characterization of 3D Localization Techniques for Particle-Tracking and Super-Resolution Microscopy
Optics Express, 17(10):8264-8277 (2009)
Three-dimensional (3D) particle localization at the nanometer scale plays a central role in 3D particle tracking and 3D localization-based super-resolution microscopy. Here we introduce a localization algorithm that is independent of theoretical models and therefore generally applicable to a large number of experimental realizations. Applying this algorithm and a convertible experimental setup we compare the performance of the two major 3D techniques based on astigmatic distortions and on multiplane detection. In both methods we obtain experimental 3D localization accuracies in agreement with theoretical predictions and characterize the depth dependence of the localization accuracy in detail.
Full text
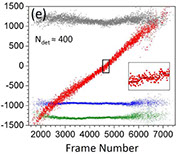
M. F. Juette, T. J. Gould, M. D. Lessard, M. J. Mlodzianoski, B. S. Nagpure, B. T. Bennett, S. T. Hess, J. Bewersdorf
Three-dimensional sub-100 nm Resolution Fluorescence Microscopy of Thick Samples
Nature Methods, 5(6):527-529 (2008)
Imaging volumes as thick as whole cells at three-dimensional (3D) super-resolution is required to reveal unknown features of cellular organization. We report a light microscope that generates images with translationally invariant 30 30 75nm resolution over a depth of several micrometers. This method, named biplane (BP) FPALM, combines a double-plane detection scheme with fluorescence photoactivation localization microscopy (FPALM) enabling 3D sub-diffraction resolution without compromising speed or sensitivity.
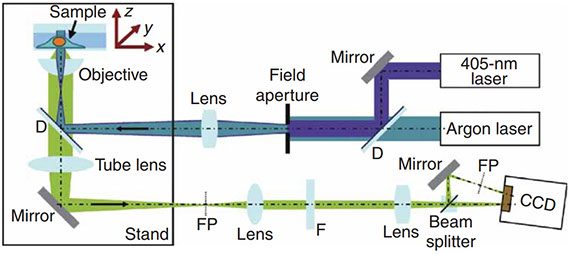
Full text
J. Bewersdorf, B. T. Bennett, K. L. Knight
Novel H2AX Chromatin Structures Revealed by 4Pi Microscopy
Proceedings of the National Academy of Sciences, 103: 18137-18142 (2006)
DNA double-strand breaks (DSBs) caused by cellular exposure to genotoxic agents or produced by inherent metabolic processes initiate a rapid and highly coordinated series of molecular events resulting in DNA damage signaling and repair. Phosphorylation of histone H2AX to form γ-H2AX is one of the earliest of these events and is important for coordination of signaling and repair activities. An intriguing aspect of H2AXphosphorylation is that γ-H2AX spreads a limited distance up to 1–2 Mbp from the site of a DNA break in mammalian cells. However, neither the distribution of H2AX throughout the genome nor the mechanism that defines the boundary of γ-H2AX spreading have yet been described. Here, we report the identification of previously undescribed H2AX chromatin structures by successfully applying 4Pi microscopy to visualize endogenous nuclear proteins. Our observations suggest that H2AX is not distributed randomly throughout bulk chromatin, rather it exists in distinct clusters that themselves are uniformly distributed within the nuclear volume. These data support a model in which the size and distribution of H2AX clusters define the boundaries of γ-H2AX spreading and also may provide a platform for the immediate and robust response observed after DNA damage.

Full text
R. Medda, S. Jakobs, S. W. Hell, J. Bewersdorf
Enhanced high-resolution imaging with quantum dots in 4Pi microscopy
Journal of Structural biology, 156: 517–523 (2006)
Imaging volumes as thick as whole cells at three-dimensional (3D) super-resolution is required to reveal unknown features of cellular organization. We report a light microscope that generates images with translationally invariant 30 30 75nm resolution over a depth of several micrometers. This method, named biplane (BP) FPALM, combines a double-plane detection scheme with fluorescence photoactivation localization microscopy (FPALM) enabling 3D sub-diffraction resolution without compromising speed or sensitivity.
Full text
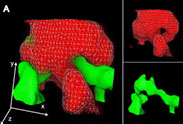

J. Bewersdorf, R. Schmidt and S. W. Hell
Comparison of I5M and 4Pi-microscopy
Journal of Microscopy, 222: 105-117 (2006)
The axial (z-) resolution of ∼100 nm provided by 4Pi and I5M fluorescence microscopy relies on the coherent addition of spherical wavefronts of two opposing high aperture angle lenses. Both microscopes feature a point-spread function (PSF) with a sharp central spot that is accompanied by axially shifted sidelobes which leads to replication artefacts in the raw image data. In a 4Pi-microscope the sidelobes are less pronounced than in I5M and without relevant lateral (x,y) substructure, making their posterior removal in the image reliable and fast. On the other hand, high speeds of raw data acquisition are more easily gained by I5M. Moreover, I5M features a stronger signal as compared to the commonly employed two-photon excitation (2PE) 4Pi-imaging mode. We investigate here the capability of both techniques to image (aqueous) specimens without artefacts. To this end, we consider the optical transfer function (OTF) of the two microscopes in conjunction with the signal-to-noise-ratio (SNR) of the object to be imaged. The imaging of E. coli bacteria with an interconvertable setup enabled a direct comparison of the two imaging modes. As both systems rely on high aperture angles, water-immersion lenses of the largest numerical aperture available (NA = 1.2) were employed. The experimental results are corroborated by simulations assuming the signal strength encountered in the experiment. The comparison of the theoretical with the experimental PSFs/OTFs showed that our setup operated close to theory in both imaging modes. Although I5M provided about 10 times brighter raw image data as compared to (2PE) 4Pi-microscopy, the I5M data could not be entirely cleared of artefacts. In conclusion, with the current aperture angles and fluorescence signal strengths, it is not advisable to trade in the suppression of the sidelobes for a larger image signal.
Full text
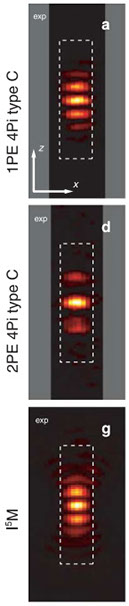

J. Bewersdorf, A. Egner, S. W. Hell
“4Pi Microscopy” and “Multifocal Multi-Photon Microscopy” in Handbook of Biological Confocal Microscopy
Pawley, J. (Ed.), 3rd ed., Springer (2006)
This third edition of a classic text in biological microscopy includes detailed descriptions and in-depth comparisons of parts of the microscope itself, digital aspects of data acquisition and properties of fluorescent dyes, the techniques of 3D specimen preparation and the fundamental limitations, and practical complexities of quantitative confocal fluorescence imaging.
Full text

H. Gugel*, J. Bewersdorf*, S. Jakobs, J. Engelhardt, R. Storz and S. W. Hell
Cooperative 4Pi excitation and detection yields 7-fold sharper optical sections in live cell microscopy
Biophysical Journal, 87: 4146-4152 (2004) * equal authorship
Although the addition of just the excitation light field at the focus, or of just the fluorescence field at the detector is sufficient for a three- to fivefold resolution increase in 4Pi-fluorescence microscopy, substantial improvements of its optical properties are achieved by exploiting both effects simultaneously. They encompass not only an additional expansion of the optical bandwidth, but also an amplified transfer of the newly gained spatial frequencies to the image. Here we report on the realization and the imaging properties of this 4Pi microscopy mode of type C that also is the far-field microscope with the hitherto largest aperture. We show that in conjunction with two-photon excitation, the resulting optical transfer function displays a sevenfold improvement of axial three-dimensional resolution over confocal microscopy in aqueous samples, and more importantly, a marked transfer of all frequencies within its inner region of support. The latter is present also without the confocal pinhole. Thus, linear image deconvolution is possible both for confocalized and nonconfocalized live-cell 4Pi imaging. Realized in a state-of-the-art scanning microscope, this approach enables robust three-dimensional imaging of fixed and live cells at ∼80nm axial resolution.
Full text
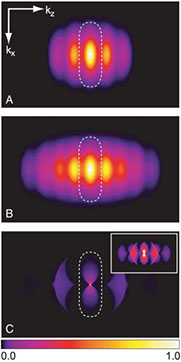
J. Bewersdorf, R. Pick and S. W. Hell
Multifocal Multiphoton Microscopy
Optics Letters, 23(9): 655-657 (1998)
We present a real-time, direct-view multiphoton excitation fluorescence microscope that provides three-dimensional imaging at high resolution. Using a rotating microlens disk, we split the near-infrared light of a mode-locked titanium–sapphire laser into an array of beams that are transformed into an array of high-aperture foci at the object. We typically scan at 225 frames per second and image the fluorescence with a camera that reads out the images at video rate. For 1.4 aperture oil and 1.2 water immersion lenses at 780-nm excitation we obtained axial resolutions of 0.84 and 1.4μm , respectively, which are similar to that of a single-beam two-photon microscope. Compared with the latter setup, our system represents a 40–100-fold increase in efficiency, or imaging speed. Moreover, it permits the observation with the eye of high-resolution two-photon images of (live) samples.
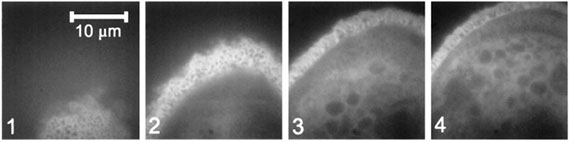
Full text
J. Bewersdorf and S. W. Hell
Picosecond pulsed two-photon imaging with repetition rates of 200 and 400 MHz
Journal of Microscopy, 191: 28-38 (1998)
We show the viability of high-resolution two-photon fluorescence imaging of fixed and live cells by exciting the fluorophores with a train of near-infrared pulses with duration in the picosecond range. This is exemplified with a compact, diode-pumped Nd:YVO4 laser, emitting trains of 7-ps pulses at a wavelength of 1064 nm, with a repetition rate of 200 MHz at two separate outputs. Incoherent combination of the outputs enabled two-photon excitation with a repetition rate of 400 MHz. For a numerical aperture of 1.4 (oil), we used an average illumination power of up to 20–40 mW at the sample. The pulses were coupled into a beam scanning microscope, either directly or through a single mode glass fibre. Compared with standard femtosecond titanium–sapphire excitation conditions, our experiments were performed with a 2.5 or 5 times higher repetition rate, 30–70 times longer pulses and 10–35 times lower pulse peak intensity. The experiments indicate the possibility of significantly relaxing the temporal pulse width constraints for a series of applications.
Full text